Primary thrombotic microangiopathy (TMA) syndromes encompass diseases that present with thrombosis in small and medium sized blood vessels due to endothelial injury. They are specific disorders that require specific treatment. The initial assessment is focused on confirming that the patient has true microangiopathic hemolytic anemia (MAHA) with or without thrombocytopenia. If MAHA and thrombocytopenia are confirmed it is important to differentiate the primary etiologies, which include:
- Thrombotic thrombocytopenic purpura, or TTP, which occurs as a result of severe ADAMTS13 deficiency
- Atypical Hemolytic uremic syndrome, or aHUS, which occurs as a result of complement dysregulation
- Hemolytic uremic syndrome, or HUS, which occurs as a result of Shiga toxins
TTP results from a severe deficiency of ADAMTS13 (defined as activity <5-10%). ADAMTS13 (a disintegrin and metalloproteinase with a thrombospondin type 1 motif, member 13) is an enzyme that cleaves large von Willebrand multimers. TTP can be hereditary (called Upshaw-Shulman syndrome) but is usually acquired as a result of an inhibitory autoantibody toward ADAMTS13. The decision to treat TTP is usually clinical since ADAMTS13 assays are often not available and may take several days to return a result if sent to a reference laboratory. TTP typically has more systemic manifestations of organ injury compared to other primary TMA syndromes.
Atypical HUS (aHUS) occurs via a complement-mediated pathway most commonly due to gene mutations of complement factors. Patients with antibodies to complement proteins comprise a smaller proportion of aHUS patients. Penetrance of aHUS is low with only a fraction of gene mutation carriers developing the syndrome. In most cases, a complement activation trigger precedes the manifestation of aHUS.
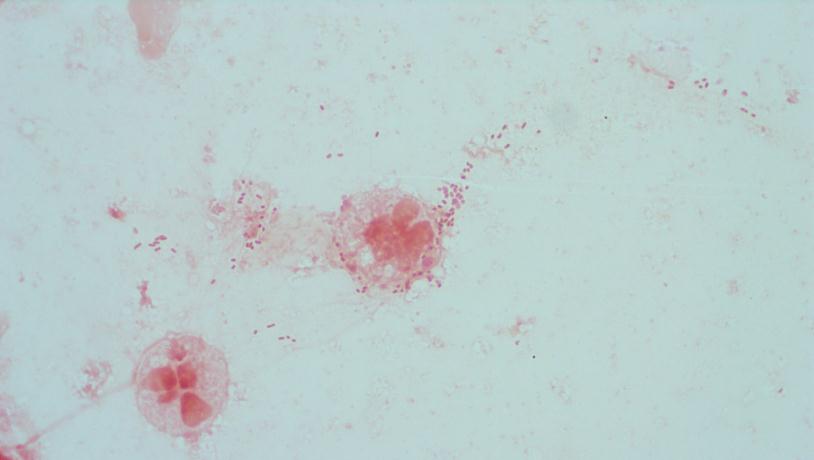
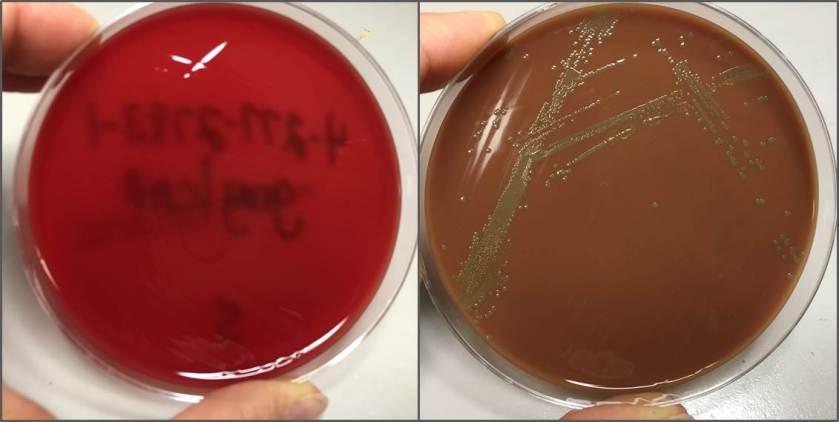
Most cases of HUS are sporadic, resulting from Shiga toxins produced by Shigella dysenteriae and some serotypes of Escherichia coli (especially O157:H7 and O104:H4). Shiga toxins preferentially injure the renal system by binding to CD77 on kidney epithelial and mesangial cells and endothelial cells. This binding causes downstream ribosomal inactivation leading to programmed cell death (apoptosis). The sporadic form of HUS is associated with bloody diarrhea.
The rationale for focusing on the distinction between HUS, aHUS and TTP is due to the different treatment options for each disease. TTP requires urgent plasma exchange to prevent death, while HUS requires treatment of the infection with little benefit from plasma exchange, while aHUS requires treatment with specialized anti-complement medication. Plasma exchange has inherent risks, so determining the cause of TMA is crucial. While no single clinical feature can be used to determine whether TTP, aHUS, or HUS is responsible for a patient’s symptoms, distinguishing factors include:
- Patient age – any primary TMA syndromes may occur at any age; however, children typically present with HUS, aHUS, or hereditary TTP, while adults more commonly present with acquired TTP
- Kidney injury –the degree of injury is usually less in TTP than in HUS; aHUS affects the arterioles while HUS tends to affect the glomeruli
- Systemic symptoms – up to two-thirds of TTP patients will have some neurologic symptoms; overtly bloody diarrhea is more typical for HUS; aHUS primarily involves kidney injury
- A previous episode or family history of aHUS – indication for screening for complement dysregulation in aHUS
Want to learn more? Check out: George JN, Nester CM. Syndromes of thrombotic microangiopathy. N Engl J Med. 2014;371(7):654-66.

-Thomas S. Rogers, DO is a third-year resident at the University of Vermont Medical Center, a clinical instructor at the University of Vermont College of Medicine, and the assistant medical director of the Blood Bank and Transfusion Medicine service.