Case History
A 36 year old Caucasian male presented to the hospital with symptoms of a bowel obstruction. His past medical history was significant for a gunshot wound to the abdomen followed by the development of colon cancer with metastasis to the liver. Recently, he had an intestinal stent placed in order to relieve an obstruction from tumoral growth. During the current admission, he was taken to the operating room for a diverting colostomy and two days later developed a fever of 101.1°F and increasing abdominal pain. Due to a concern for sepsis, blood cultures were collected and piperacillin-tazobactam was added to his antibiotic therapy regimen which already included vancomycin and ciprofloxacin.
Laboratory Identification


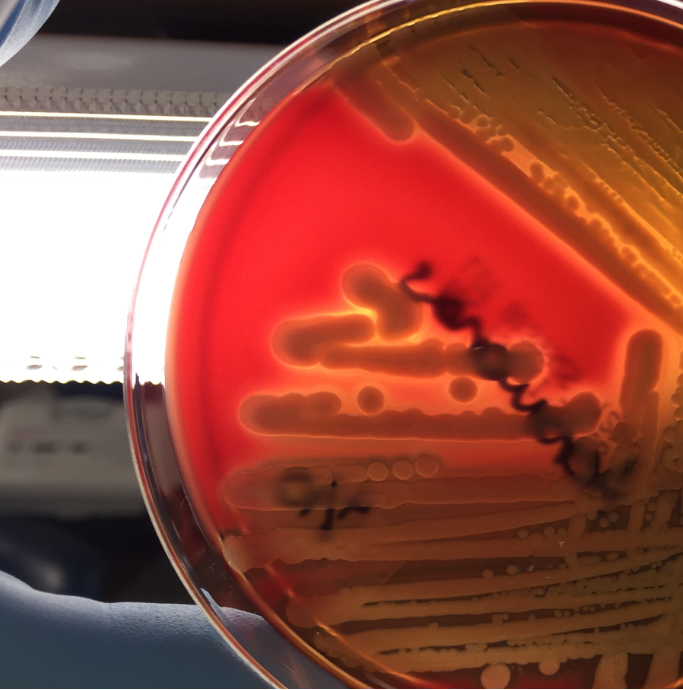
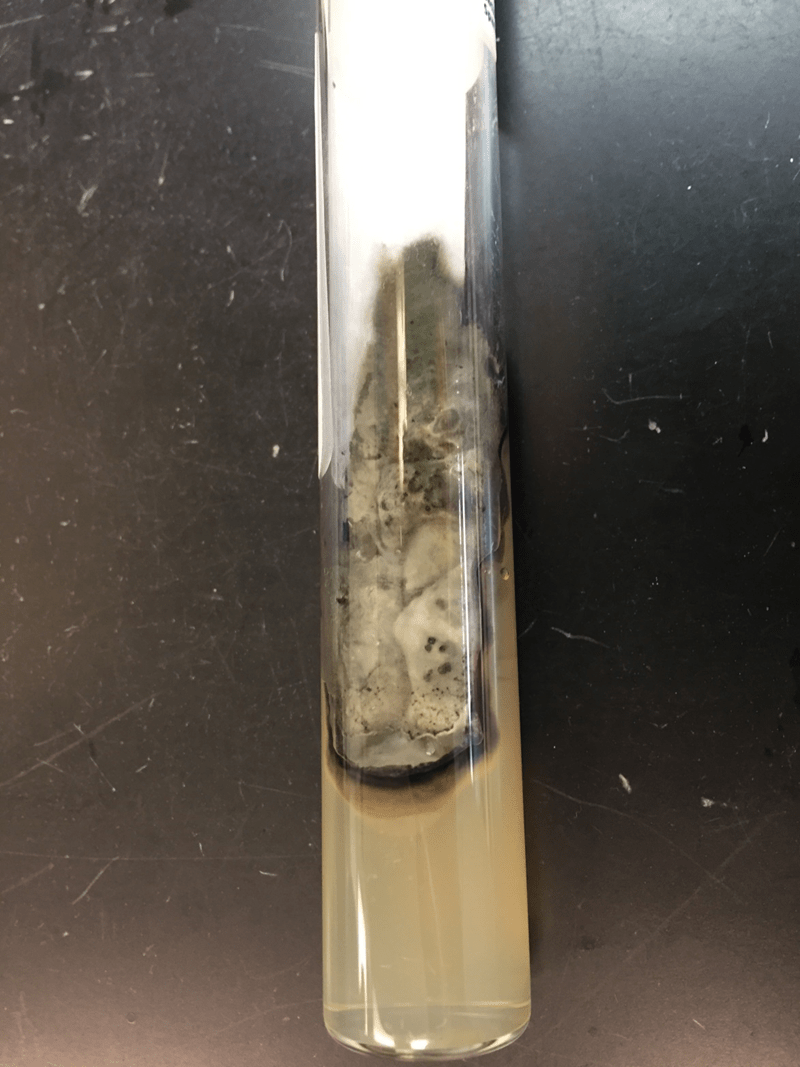
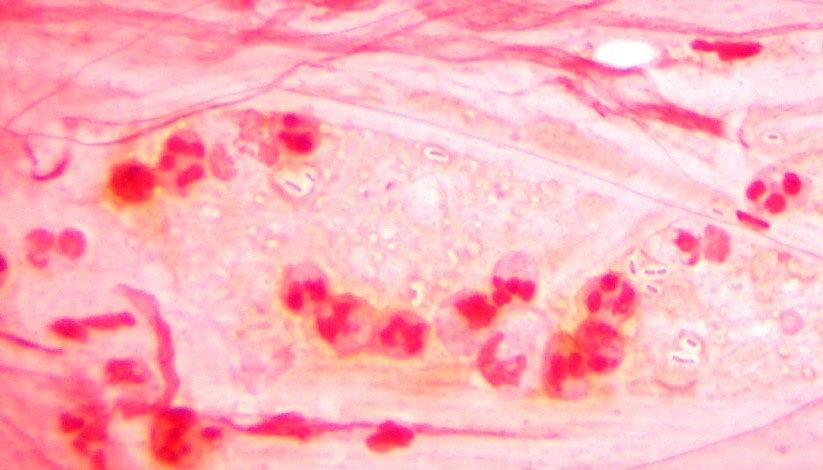
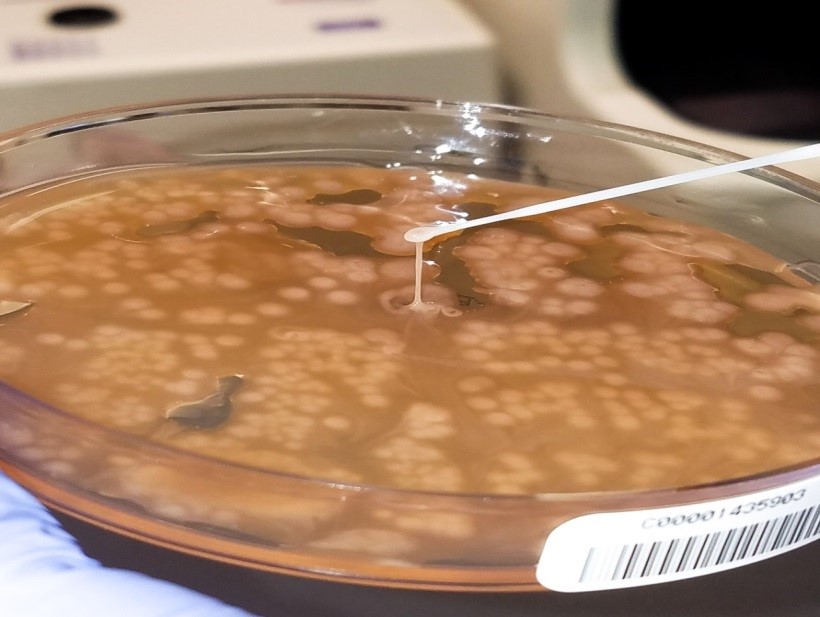
The blood culture signaled positive after less than 24 hours on the automated instrument and Gram stain revealed gram negative rods (Image 1). The organism grew as mucoid colonies on blood, chocolate and MacConkey agars after 24 hours of incubation at 35°C in 5% CO2 (Image 2). When the blood agar plate was examined with a light source, the organism clearly illustrated beta hemolysis (Image 3). Rapid biochemical testing was positive for catalase, oxidase and indole. The Vitek II identified the isolate as Aeromonas veronii bv. sobria.
Discussion
Aeromonas spp. are facultative anaerobic gram negative rods. They are inhabitants of aquatic ecosystems and as such can cause wound infections in people exposed to polluted lakes or brackish water with fresh breaks in their skin. Additionally, gastroenteritis is common with Aeromonas spp. and are often acquired through ingestion of unpurified water or to a lesser extent by consumption of contaminated meats, fresh produce or dairy products. Extraintestinal infections, including sepsis and meningitis, can result by spread from GI sources or wound infections. Interesting, medicinal leeches, used in the post-operative setting to increase blood flow to the surgical site, are colonized with Aeromonas spp. (most commonly Aeromonas veronii bv. sobria) and can result in systemic infections in the patient.
In the laboratory, Aeromonas spp. grow readily from stool, wound and blood sources on commonly used media and exhibits beta hemolysis on blood agar. In addition, Aeromonas spp. will grow on CIN agar (at room temperature as well as incubator temperature) as colonies with a pink center surrounded by a white apron and are indistinguishable from Yersinia spp. Aeromonas spp. is positive for catalase, oxidase and indole by rapid testing. In most cases, identification of Aeromonas spp. to the complex level can be accomplished by biochemical testing (esculin, VP), automated instrumentation or MALDI-TOF mass spectrometry. The three clinically relevant complexes include: A. hydrophila complex, A. caviae complex and A. veronii complex.
With regards to susceptibility testing for Aeromonas spp., the CLSI M45, 3rd edition provides guidelines for the three complexes discussed above. Third or fourth generation cephalosporins, fluoroquinolones and trimethoprim-sulfamethoxazole are recommended as antibiotics for primary testing for isolates from extraintestinal sites. Aeromonas spp. are uniformly resistant to ampicillin, amoxicillin-clavulanate and cefazolin and many strains may possess various inducible beta lactamases.
In the case of our patient, with the laboratory identification of Aeromonas veronii bv. sobria, his gram negative coverage was switched to ciprofloxacin for a 10 day course. His PICC was removed and on further imaging studies he was found to have a large abdominal abscess which required surgical drainage. Subsequent cultures from the abdominal abscess were negative.

-Lisa Stempak, MD, is an Assistant Professor of Pathology at the University of Mississippi Medical Center in Jackson, MS. She is certified by the American Board of Pathology in Anatomic and Clinical Pathology as well as Medical Microbiology. She is the director of the Microbiology and Serology Laboratories. Her interests include infectious disease histology, process and quality improvement, and resident education.