This is not your Mom’s PCR. These new kids on the block are making PCR extremely fast. PCR (Polymerase Chain Reaction) technology won the Nobel Prize for allowing molecular research to advance much more rapidly (for an interesting read on the quirky Laureate who gave up science to go surfing, read more here: Wikipedia ). It has become the most commonly used work horse of most molecular diagnostic assays, usually in the form of real-time PCR. It is used for a variety of purposes from detecting bacteria and viruses, identity testing for forensics and bone marrow engraftment, cancer mutation analysis, and even sequencing by synthesis used by Illumina for massively parallel sequencing.
This technique is still limited by requiring highly trained technologists to perform DNA extraction, time-consuming processing, and the time of real-time PCR itself. Overall, this process takes about a 5-8 hours. While this is much faster than in the past, it would be unacceptable for use in the point-of-care (POC).
But why would DNA testing need to be POC? The term sounds like an oxymoron in a field where many results have a 2-month turnaround time. There are certain circumstances where molecular testing would impact patient care. For instance, a doctor testing a patient in their office for a sexually transmitted infection would want to know if they have gonorrhea/ chlamydia so they could prescribe proper antibiotics. Similarly, POC molecular testing could be applied in a bioterrorism incident to test samples for an infectious agent. Or POC testing would benefit low-resource areas internationally where HIV testing could be used to manage anti-retroviral therapy in patients many miles from a laboratory.
For PCR as a test to be useful at the POC setting, it would have to provide a result within 10-15 minutes and be performed as a waived test. Two recent examples the demonstrate how this is possible have been highlighted at recent conferences of the American Association of Clinical Chemistry, which I just got back from: Extreme PCR1 and Laser-PCR.2
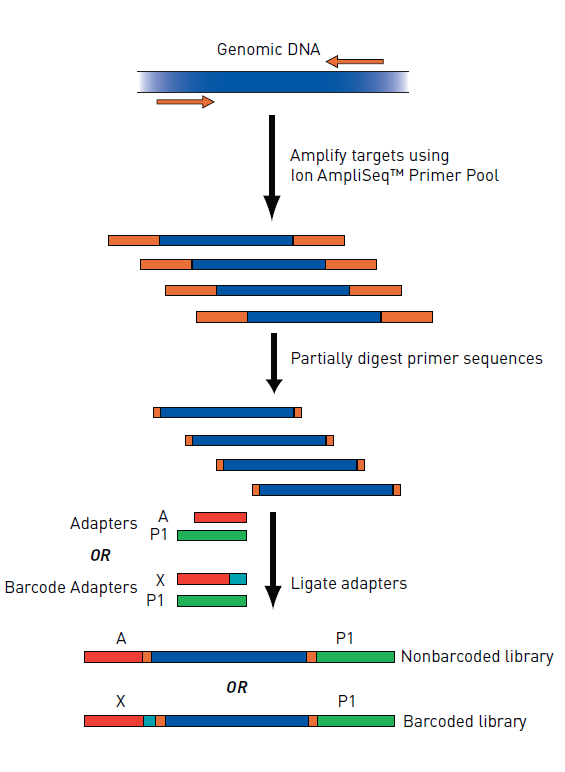
Extreme PCR refers to a technique of rapidly cycling the temperature of PCR reactions. The reaction occurs in a thin slide that evenly distributes the reagents, temperature and is clear to permit easy reading of fluorescence measurements (Figure 1). DNA Polymerase enzyme and primers to amplify the target DNA are added at much higher concentrations than normal (20x).

This flies in the face of traditional PCR chemistry dogma as specificity would plummet and normal DNA could be amplified instead of target DNA. This would create a false positive. However, let’s think about what is actually happening with non-specific reactions. Primers are designed to match one region of DNA, which is very unique within the whole genome. However, the genome is so large that some segment may look very similar and be different in just 1 or 2 of the 20 base pairs that a primer matches. A primer could bind to this alternate region but less efficiently. So, the binding would be weaker and take more time to occur.
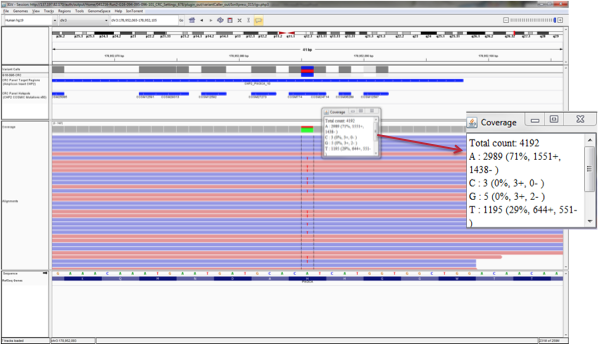
Therefore, by speeding up the cycling time to just a few seconds, only the most specific interactions can take place and non-specific binding is offset (Figure 2)!

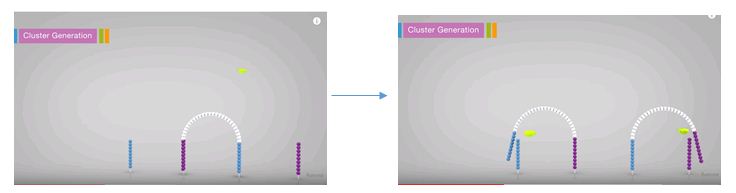
Laser PCR does not report the use of increased reagents like Extreme PCR (it may be proprietary), but they boast a very innovative method to quickly heat and cool PCR reactions. GNA Biosciences use gold nanoparticles with many DNA adapters attached (Watch the video below for a great visual explanation!).
These adapters are short sequences of DNA that bring the target DNA and primers together to amplify the target DNA sequence. Then as the name implies, a laser zaps the gold beads and heats them up in a very localized area that releases the DNA strands. The released DNA binds another gold particle, replicates, rinses, and repeats. The laser energy thus heats the gold in a small area that allows for quick heating and cooling within a matter of seconds.
These new PCR methods are very interesting and can have a big impact on changing how molecular pathology advances are brought to the patient. On a scientific note, I hope you found them as fascinating as I did!
References
- Myrick JT, Pryor RJ, Palais RA, Ison SJ, Sanford L, Dwight ZL, et al. Integrated extreme real-time PCR and high-speed melting analysis in 52 to 87 seconds. Clin Chem 2019;65:263–71.
- CLN Stat. A Celebration of Innovation. AACC’s first disruptive technology award to recognize three breakthrough diagnostics. https://www.aacc.org/publications/cln/cln-stat/2018/july/10/a-celebration-of-innovation
- G. Mike Makrigiorgos. Extreme PCR Meets High-Speed Melting: A Step Closer to Molecular Diagnostics “While You Wait” Clin Chem 2019.

-Jeff SoRelle, MD is a Chief Resident of Pathology at the University of Texas Southwestern Medical Center in Dallas, TX. His clinical research interests include understanding how the lab intersects with transgender healthcare and improving genetic variant interpretation.