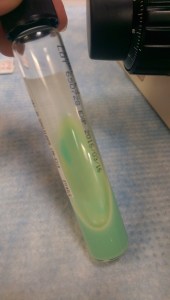
A cornerstone of good clinical microbiology laboratory practice is to look for visual clues in how organisms grow in culture. This can help quickly signal to the laboratorian that a particularly meaningful pathogen is present.
For example, a wound culture where an anaerobic blood agar plate is showing a double zone of hemolysis? The tech should immediately think that Clostridium perfrigens may be present. An abscess culture growing a ridiculously mucoid colony of a lactose-fermenting Gram negative rod? Hyper-mucoid Klebsiella pneumoniae is a hypervirulent strain associated with abscess formation. Training and experience in what to look for in cultures is one of the fascinating and exciting (and potentially daunting) aspects of clinical microbiology!
But crucially, sometimes the textbook visual association is NOT as specific a finding as we may believe! One such example: Mycobacterium species and cording.
Cording and M. tuberculosis
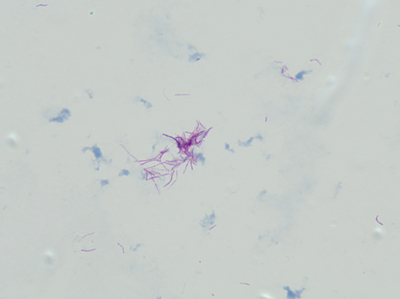
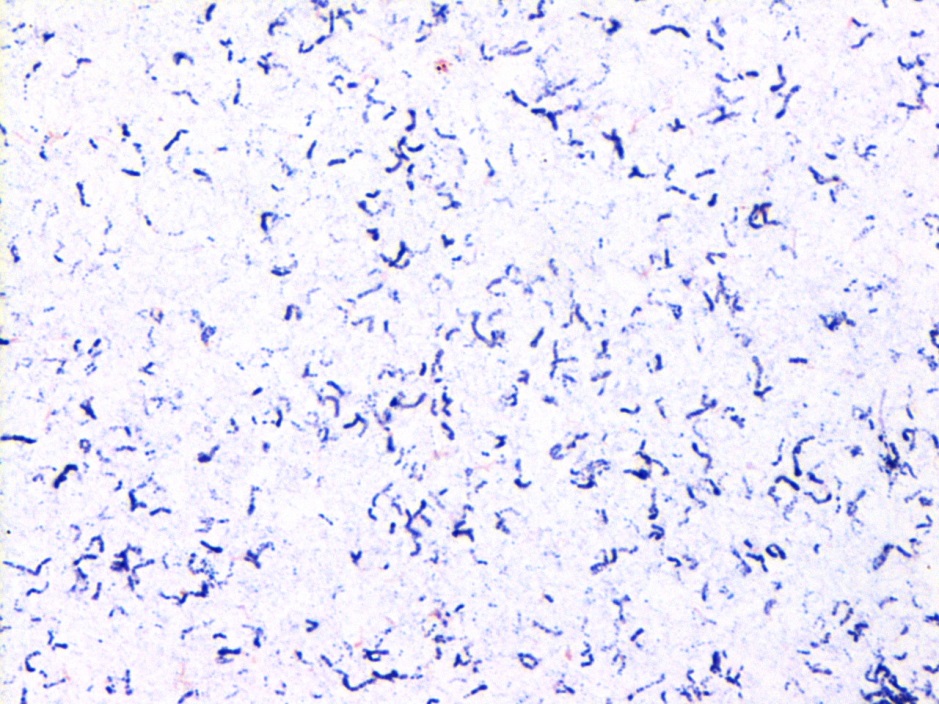
“Cording” is term used to describe twisting, serpentine appearance of Mycobacterium in liquid broth culture. And, at least in many places, it is taught to budding microbiologists and infectious disease clinicians as a hallmark characteristic of Mycobacterium tuberculosis (Mtb).

How long have people been observing Mtb cording in liquid culture? From very beginning!
Robert Koch, originator of the famous Koch’s Postulates, used Mtb (now recognized to be just one member of the M. tuberculosis complex) to demonstrate that disease was caused by discrete organisms originating from one host and infecting another, what we accept as germ theory. In his original description of Mtb, Koch wrote that the bacilli would “ordinarily form small groups of cells which are pressed together and arranged in bundles.”
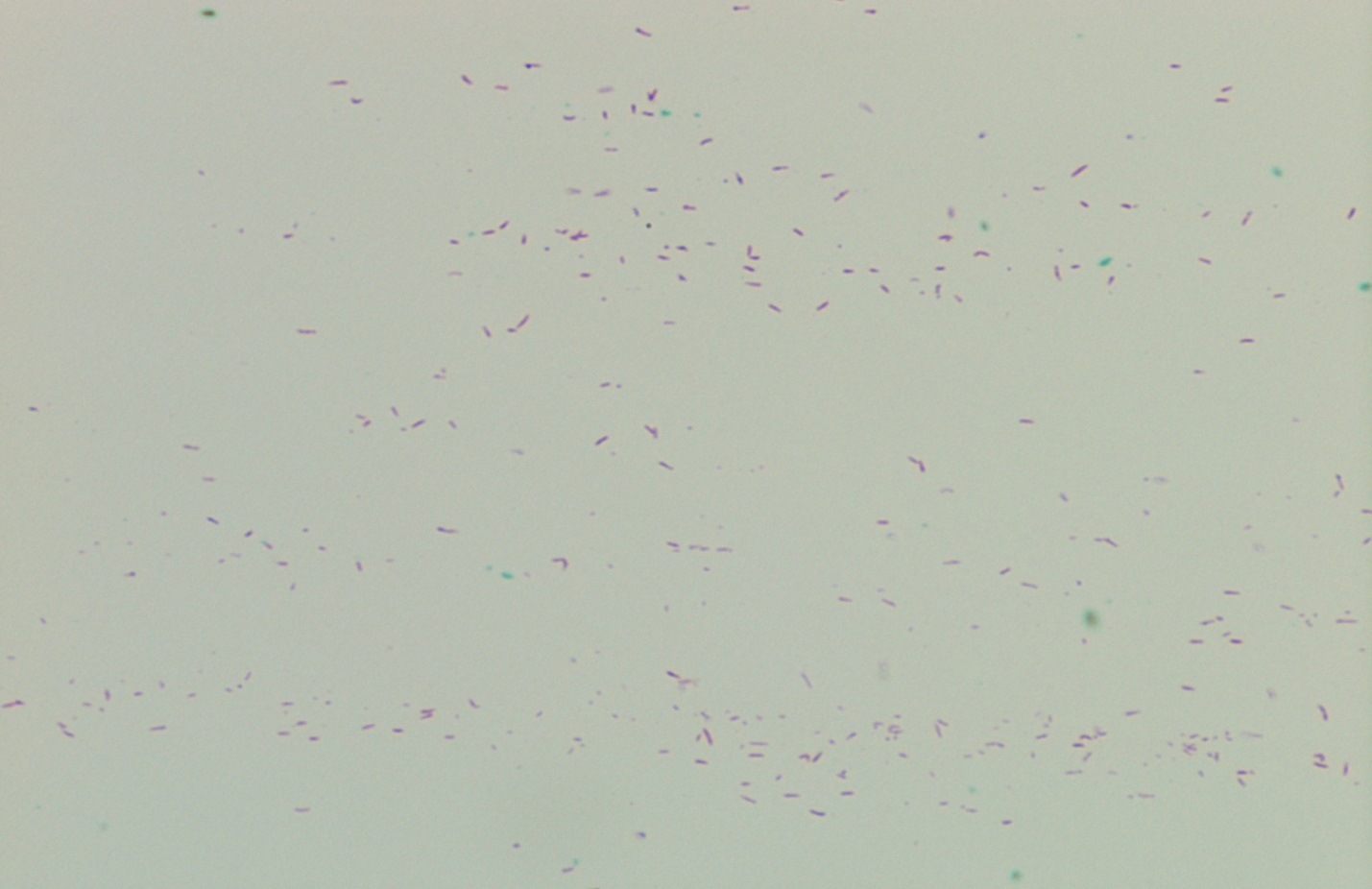
Cording phenotype is distinct from “clumping.” With cording, bacilli lay in tightly packed parallel strands, and not clumped together showing random bacilli orientation.
The phenomenon of cording is distinct to organisms cultured in liquid media like Middlebrook 7H9 broth. But in 1947 Middlebrook (where Middlebrook 7H9 broth gets its name), Dubos and Pierce published a key paper showing that cording Mtb strains also grew as rough colonies on solid agar and that this phenotype was associated with virulent and not avirulent Mtb strains.
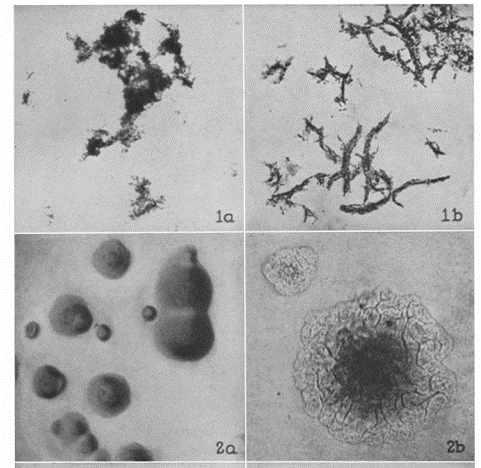
Only a few years later in 1950, the actual cell wall component, the so-called “cording factor,” that causes these growth characteristics was extracted and identified. Trehalose-6,6-dimycolate (TDM), a major glycolipid from the cell walls of virulent strains of Mtb.
Further work suggested that, not only was this TDM glycolipid the cause of the unique cording appearance and rough colonies in in vitro culture conditions, but that directly was a virulence factor: studies in animal models showed that TDM not only directly allowed Mtb to avoid phagocytosis from macrophages by virtue of forming large clusters of bacilli, but it also directly prevented macrophage intracellular killing mechanisms (see Hunter, 2006).
The association between cording and clinical isolates of Mtb was seen as a very sensitive and specific finding. Such a robust feature, in fact, that a number of publications from clinical microbiology labs specifically to cording as a reliable method to quickly identify Mtb (see McCarter 1998) or at least to select which isolates should be further identified by more specific methods(see Nelson, 1998).
Cording in non-tuberculosis Mycobacteria
Though there was a strong the association between cording and Mtb, scientists had known for years that other Mycobacterium species, outside the M. tuberculosis complex, also expressed TDM. But beginning with a case report in 2008, an isolate of Mycobacterium marinum was seen to have a biofilm with “cording morphology.” Other reports of cording in M. marinum followed (see Staropoli 2008).
Importantly, cording in non-tuberculosis Mycobacterium species is 1) actual cording and 2) visually indistinguishable from cording seen in Mtb.
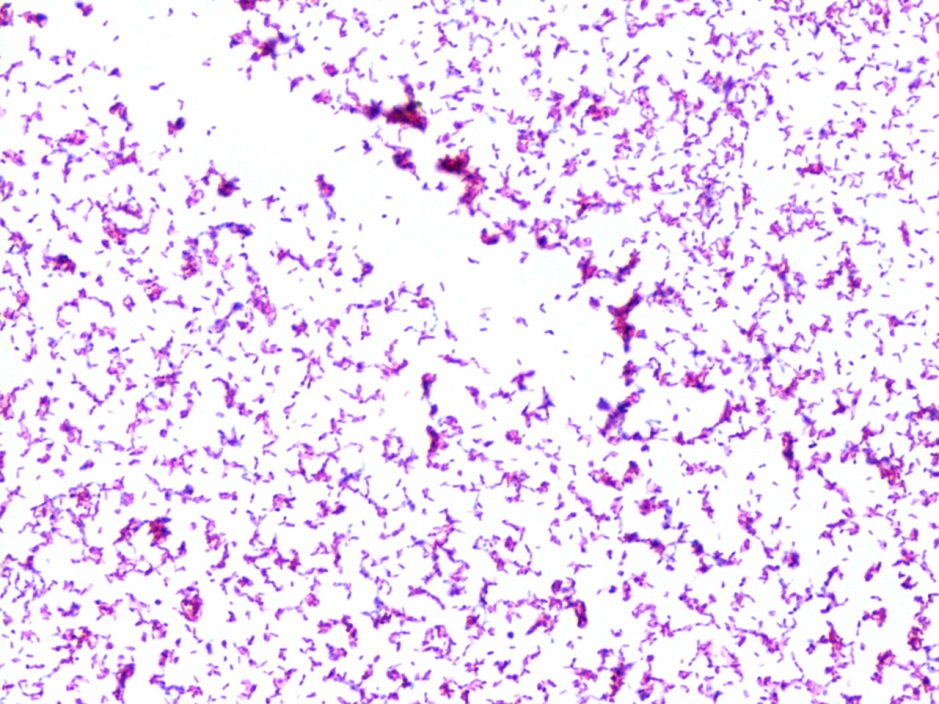
In my laboratory, (Spokane, Washington, USA) we regularly see isolates of Mycobacterium abscessus, a rapid growing mycobacterium species, with clear cording morphology. These twisting structure are clearly cords, not just clumps.

Researchers have since identified not just different levels of TDM in isolates of M. abscessus but also that those levels correlate to the known rough or smooth appearance of these isolates (see Llorens-Fons, 2017). It is interesting to consider that more virulent isolates of M. abscessus are likely to be rough, cording-type growers, similar to what is seen with Mtb.

Beyond what I hope was an interesting look into the history of Mycobacterium identification, and a great chance to show some beautiful AFB cording, it’s a good reminder that in our lab culture (pun definitely intended!) visual clues to organism identification may not fit what you were first taught.

References
- Middlebrook G, Dubos RJ, Pierce C. VIRULENCE AND MORPHOLOGICAL CHARACTERISTICS OF MAMMALIAN TUBERCLE BACILLI. J Exp Med. 1947 Jul 31;86(2):175-84. doi: 10.1084/jem.86.2.175. PMID: 19871665; PMCID: PMC2135722.
- Hunter RL, Olsen MR, Jagannath C, Actor JK. Multiple roles of cord factor in the pathogenesis of primary, secondary, and cavitary tuberculosis, including a revised description of the pathology of secondary disease. Ann Clin Lab Sci. 2006 Autumn;36(4):371-86. PMID: 17127724.
- McCarter YS, Ratkiewicz IN, Robinson A. Cord formation in BACTEC medium is a reliable, rapid method for presumptive identification of Mycobacterium tuberculosis complex. J Clin Microbiol. 1998 Sep;36(9):2769-71. doi: 10.1128/JCM.36.9.2769-2771.1998. PMID: 9705435; PMCID: PMC105205.
- Nelson SM, Cartwright CP. Comparison of algorithms for selective use of nucleic-acid probes for identification of Mycobacterium tuberculosis from BACTEC 12B bottles. Diagn Microbiol Infect Dis. 1998 Aug;31(4):537-41. doi: 10.1016/s0732-8893(98)00049-2. PMID: 9764392.
- Hall-Stoodley L, Brun OS, Polshyna G, Barker LP. Mycobacterium marinum biofilm formation reveals cording morphology. FEMS Microbiol Lett. 2006 Apr;257(1):43-9. doi: 10.1111/j.1574-6968.2006.00143.x. PMID: 16553830.
- Staropoli JF, Branda JA. Cord formation in a clinical isolate of Mycobacterium marinum. J Clin Microbiol. 2008 Aug;46(8):2814-6. doi: 10.1128/JCM.00197-08. Epub 2008 Jun 25. PMID: 18579723; PMCID: PMC2519507.
- Llorens-Fons M, Pérez-Trujillo M, Julián E, Brambilla C, Alcaide F, Byrd TF, Luquin M. Trehalose Polyphleates, External Cell Wall Lipids in Mycobacterium abscessus, Are Associated with the Formation of Clumps with Cording Morphology, Which Have Been Associated with Virulence. Front Microbiol. 2017 Jul 25;8:1402. doi: 10.3389/fmicb.2017.01402. PMID: 28790995; PMCID: PMC5524727.

-Dr. Richard Davis, PhD, D(ABMM), MLS(ASCP)CM is a clinical microbiologist and regional director of microbiology for Providence Health Care in Eastern Washington. A certified medical laboratory scientist, he received his PhD studying the tropical parasite Leishmania. He transitioned back to laboratory medicine (though he still loves parasites!), and completed a clinical microbiology fellowship at the University of Utah/ARUP Laboratories in Utah before accepting his current position. He is a 2020 ASCP 40 Under Forty Honoree.