Case History
A 53 year old male presents to the emergency department with complaints of fatigue, night sweats, dyspnea, dry cough, and right knee swelling. He has multiple skin lesions including violaceous papules on his medial thigh, subcutaneous nodules on bilateral lower legs, and ulcerations on his right lateral leg. His past medical history is significant for psoriatic arthritis. Previously, he has taken adalimumab (Humira), etanercept (Enbrel), and golimumab (Simponi), but his current treatment regimen consists of methotrexate and prednisone. His recent travel history is significant for scuba diving in Thailand and honeymooning in the Caribbean. Dermatology was consulted and a punch biopsy was performed close to an ulcer on his right lateral leg and sent to surgical pathology. An additional biopsy specimen was sent to microbiology for bacterial, fungal and mycobacterial cultures.
Laboratory Identification
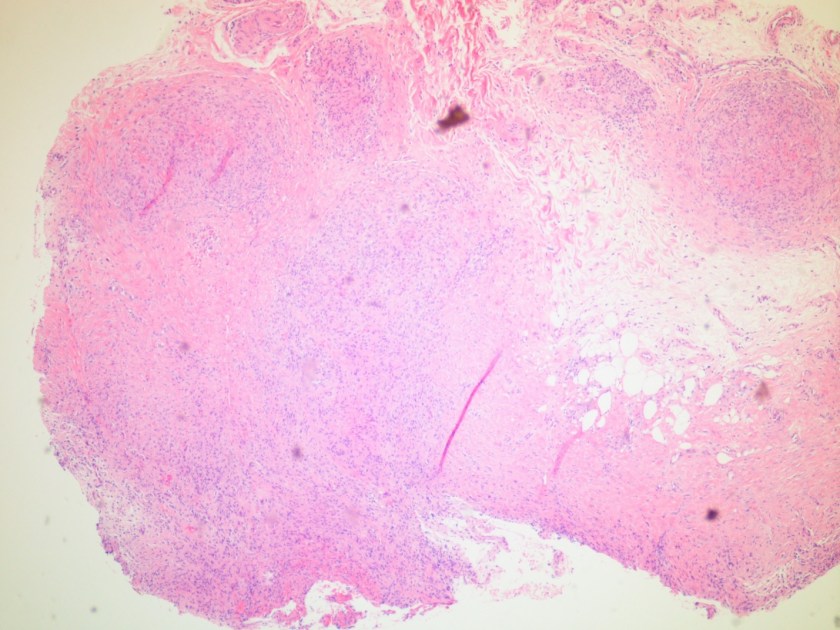
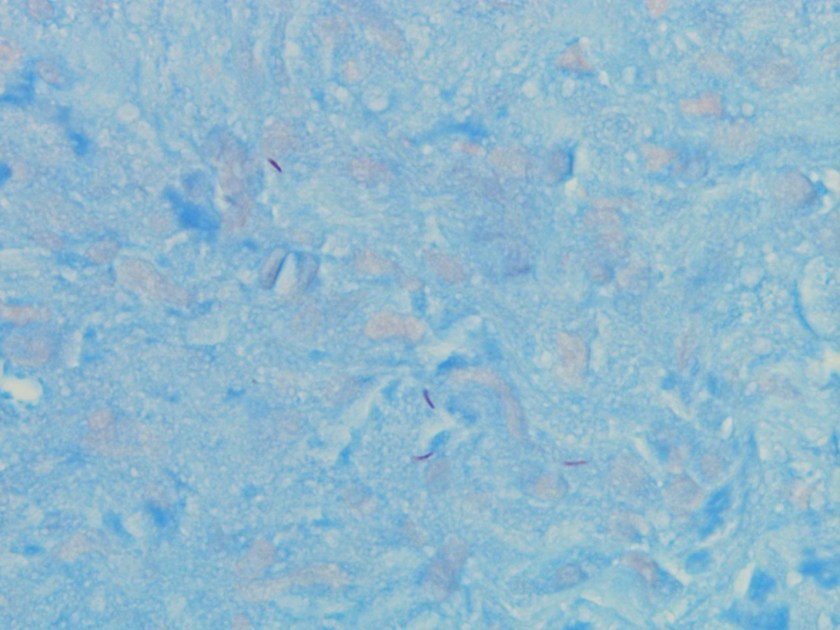
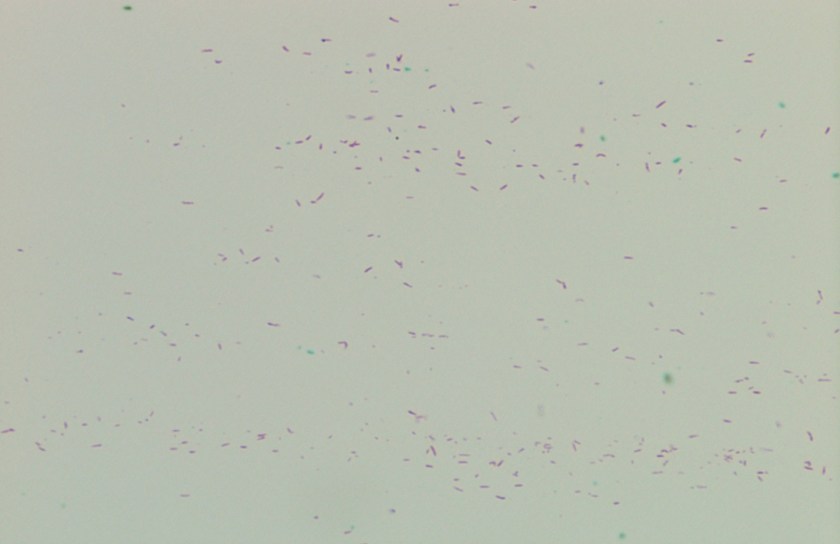
Microscopic examination of the punch biopsy skin specimen revealed a non-caseating granuloma in the deep subcutaneous tissue, with no involvement of the overlying dermis and epidermis (Figure 1). A Kinyoun stain of the tissue showed that the granuloma contained occasional acid-fast bacilli (Figure 2). The bacterial and fungal cultures sent to microbiology were negative. Portions of the specimen used to set up the mycobacterial cultures were incubated in MGITs at 32 and 37°C because the specimen source was skin. The 32°C tube, which was supplemented with hemin, gave a positive signal after 3 weeks of incubation. The Ziehl-Neelsen stain from this tube revealed numerous acid fast bacilli (Figure 3). DNA Gen-Probe analysis was negative for Mycobacterium tuberculosis complex and M. avium-intracellulare (MAI) complex. The organism was identified as M. haemophilum by pyrosequencing.
Discussion
Mycobacterium haemophilum was first identified in 1978 from an Israeli patient with Hodgkin lymphoma. It has a known predilection for infecting the skin and subcutaneous tissue in immunocompromised patients, especially those with lymphopenia as a result of acquired immune deficiency syndrome (AIDS), allogeneic bone marrow transplantation, and those on immunosuppressant therapies for rheumatologic conditions. The clinical presentation frequently consists of painful subcutaneous nodules and ulcers that can progress to abscesses and draining fistulas. Bone and joint infections have also been reported, which manifest as arthritis, tenosynovitis, and osteomyelitis. AIDS patients in particular are known to present with disseminated disease, with multiple cutaneous lesions, mainly involving the extremities. Relatively little is known about this infection and the optimal treatment is not standardized, but combinations of three or four of the following drugs have been used successfully: isoniazid, rifamycins, ciprofloxacin, amikacin, doxycycline, and clarithromycin.
Unlike the majority of mycobacteria, M. haemophilum does not grow well in culture at 37°C. Rather, it prefers lower temperatures, ideally between 28–32°C. This characteristic is shared by several other mycobacterial species that also characteristically infect the skin, including M. marinum, M. chelonae, M. abscessus, and M. ulcerans. A unique feature of M. haemophilum among the mycobacteria is that it requires hemin (X factor) to survive and will only grow in media enriched with this nutrient. Similar to Haemophilus influenzae, M. haemophilum can be cultured on chocolate agar, as well as on Middlebrook 7H10 agar incubated with an “X-factor strip” and on Lowenstein-Jensen medium containing 2% ammonium citrate.
Typically, colonies grow after 2-4 weeks of incubation at 32°C and have either a rough or smooth appearance. M. haemophilum is a non-photochromogen according to the Runyon classification system, and its colonies are buff colored and do not produce pigment in either light or dark conditions. M. haemophilum is chemically inert by traditional biochemical mycobacterial tests, with the exception of pyrazinamidase production. As illustrated by this case, DNA probe analysis is helpful with regard to the mycobacterial species it excludes, but not for speciation of less common organisms. At the present time, DNA probes exist only for M. tuberculosis complex, MAI complex, M. kansasii, and M. gordonae.
In the case of our patient, the species level identification was determined by pyrosequencing. He was treated with an extended course of 3 agents: rifabutin, clarithromycin, and moxifloxacin with good response.

–Vikas Nath, MD, is a 4th year resident in Anatomic and Clinical Pathology at the University of Mississippi Medical Center in Jackson, MS.

-Lisa Stempak, MD, is an Assistant Professor of Pathology at the University of Mississippi Medical Center in Jackson, MS. She is certified by the American Board of Pathology in Anatomic and Clinical Pathology as well as in Medical Microbiology. Currently, she oversees testing performed in both the chemistry and microbiology laboratories. Her interests include infectious disease histology, process and quality improvement, and resident education.
