One of the most common questions I’m asked by family members is “do you know when they died?” If death occurs in the hospital, or is witnessed, the time of death isn’t controversial. It’s common though in forensics that people may not be found for hours, days, weeks, or more. Forensics television shows usually depict an investigator measuring body temperature at the scene, and then confidently declaring they’ve been dead for 44 hours. Unfortunately, there aren’t any existing methods that actually give that level of precision – but there is a way we can systematically approach the question.
When determining time of death (TOD), it’s most important to keep in mind that it will be an estimate. The estimate starts with the “window of death” – the time between when the decedent was last known alive and when their body was found. The smaller this window, the greater accuracy is possible.
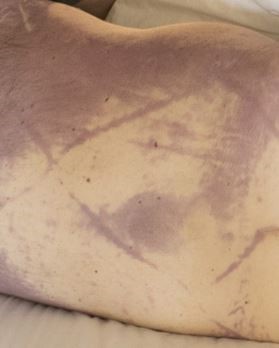
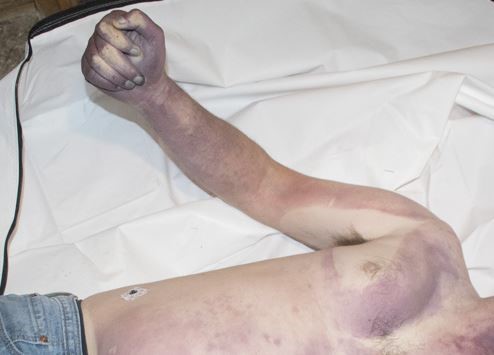
Once the window is known, one can assess postmortem changes of the body. Livor mortis is the gravity-dependent settling of blood within vessels, which can appear as soon as twenty minutes after death. Sparing of lividity will be present in areas of pressure, such as parts of the body pressed against the floor or with tight clothing. Livor is initially blanchable, but after 8 to 12 hours blood extravasates from vessels and it becomes “fixed”. Clearly though, this only allows one to differentiate between ‘less than’ or ‘greater than’ 8 to 12 hours.
Rigor mortis (stiffening of the body after death) occurs because of postmortem ATP depletion. Muscle fibers require a supply of ATP to both contract and relax – once ATP levels are sufficiently low, muscle will remain contracted until the fibers are broken down by decompositional changes. Generally speaking, rigor starts to develop within an hour of death, peaks from 12 to 24 hours, and dissipates by 36 hours. However, these are average intervals. The onset of rigor is hastened by vigorous physical activity, seizures, electrocution, or increased body temperature, which preemptively deplete ATP. Rigor is also harder to detect in people with low muscle mass (e.g. infants), and can’t be assessed in frozen bodies with those with extensive thermal damage.
Cooling of the body after death, known as algor mortis, is similarly prone to interfering elements. One can find many formulas for estimating the time of death based on the temperature of the body – unfortunately, none of them are particularly useful because of the assumptions that must be made. Change in temperature after death is affected by numerous variables, including body habitus, clothing, wind, actual body temperature at the time of death (not many people are constantly at 98.6℉), sepsis, terminal seizures, and many others. If the environment is warmer than the body, the temperature can even increase after death.
I’ll briefly mention vitreous potassium measurement, which is probably the most recently discovered (and debunked) “holy grail” of time of death. Similar to algor and rigor mortis, vitreous potassium does a reasonably decent job predicting time of death in a controlled experiment – but in this line of work, people don’t tend to die in controlled environments.
At the end of the day, time of death is best estimated by thorough scene investigation, correlated with the evidence the body provides. Newspapers or mail not retrieved from the mailbox, expiration dates on perishable groceries, last refills of prescriptions, and unreturned text messages or phone calls can all narrow down the window of death.
As stated earlier, the longer the interval between death and discovery of the body, the more difficult time of death determination becomes. In advanced decomposition, there is no rigor, livor, or algor remaining to assess (there may even be scant residual soft tissue). In one such situation, despite months of a potential “window of death”, dates on unopened bills and crossed-off calendar dates helped us place the time of death within one or two days. It’s not as flashy as multivariate equations for temperature or potassium levels, but it’s far more accurate and scientifically defensible.

-Alison Krywanczyk, MD, FASCP, is currently a Deputy Medical Examiner at the Cuyahoga County Medical Examiner’s Office.