Case History
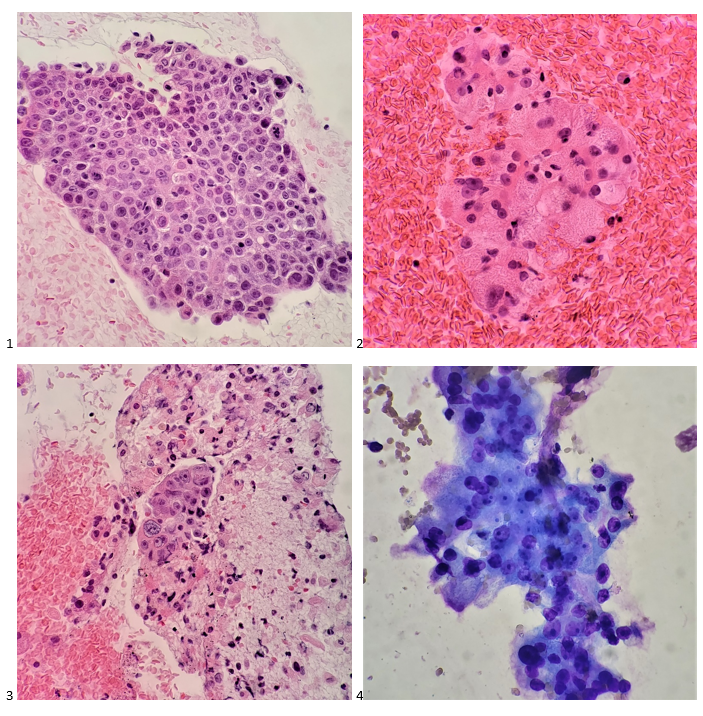
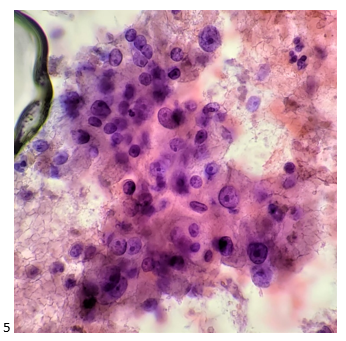
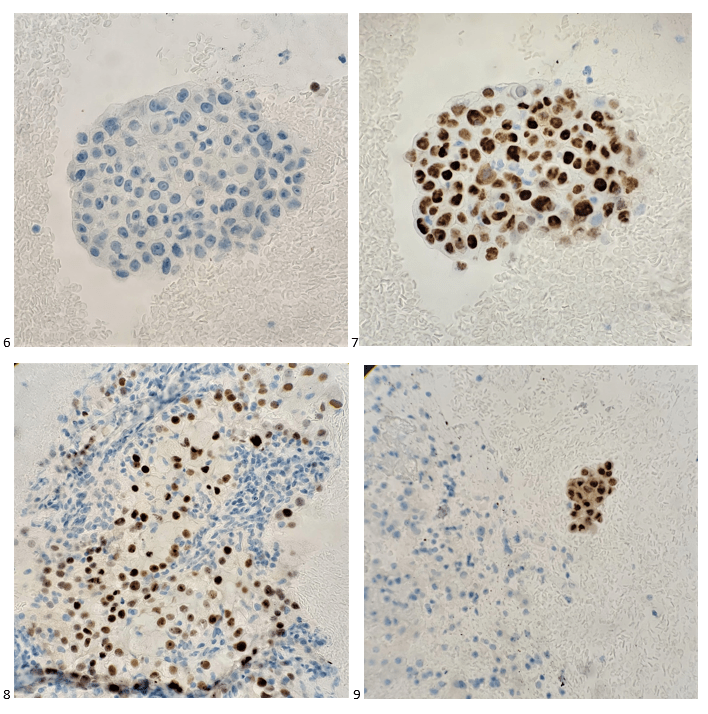
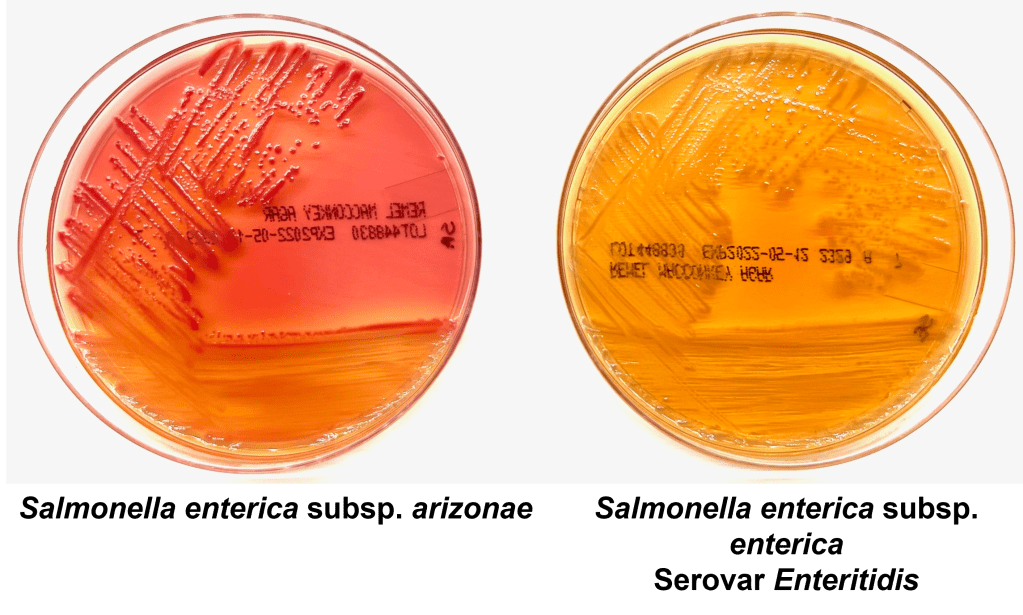
A 27 year old male presented to the Emergency Department (ED) with complaints of right knee pain and swelling for one week. Two weeks prior, he tripped while walking to work and began to feel pain in his right calf. Upon physical examination, swelling was noted in his ankles, knee, shoulders, and fingers. The knee and shoulder were tender to palpation. In the ED, he was afebrile and vitals were normal. He denied any sort of injury, chills, or rash and no history of tobacco, alcohol, or illicit substance abuse. CT scan of the lower extremity showed no acute fracture but moderate to large knee joint effusion was observed. He and his fiancé (male partner) has been in a monogamous relationship for almost a decade, however the patient did have a history of gonorrhea nine years ago but was treated. Knee arthrocentesis was performed. The fluid was yellow and cloudy and contained 27,000 WBCs. The Gram stain of the synovial fluid showed many intracellular gram negative diplococci and the joint fluid culture grew out Neisseria gonorrhoeae. PCR of the rectal swab also detected N. gonorrhoeae.
Discussion
N. gonorrhoeae is the causative agent of gonorrhea, a sexually transmitted disease. In the United States, it is the second most commonly reported communicable disease.1 While infections can be asymptomatic, in men, gonorrhea commonly causes acute urethritis with dysuria, urethral discharge, and rarely, epididymitis.2,3,4 In women, gonorrhea can cause cervicitis and lead to pelvic inflammatory disease (PID), infertility, ectopic pregnancy, and chronic pelvic pain.5,6 Those with gonococcal endocervicitis can be co-infected with Chlamydia trachomatis and/or Trichomonas vaginalis, other causative agents of sexually transmitted diseases. N. gonorrhoeae can cause extragenital infections in the pharynx and rectum, which are most commonly seen among men who have sex with men (MSM). Disseminated gonococcal infection is rare (0.5-3% of infected individuals) and can be characterized by low grade fever, hemorrhagic skin lesions, tenosynovitis, polyarthralgia and septic arthritis. Complications of disseminated infections may include permanent joint damage, endocarditis, and meningitis. Gonococcal conjunctivitis mainly affects newborns from untreated mothers.7
Gonorrhea can be diagnosed clinically by a history and physical examination and also, by microbiological methods. Home collection kits are available to increase convenience. On a Gram stain, N. gonorrhoeae, a gram negative coccus, frequently appears within or closely associated polymorphonuclear leukocytes (PMNs) typically as diplococci pairs. Direct smears can be prepared from urethral, endocervical sites, and normally sterile or minimally contaminated sites such as joint fluid. Swab specimens (e.g. urogenital, pharyngeal, vaginal or rectal) should be collected with a Dacron or Rayon swab as calcium alginate and cotton swabs may be toxic or inhibitory for the bacteria.8 Specimens must be transported to the microbiology immediately. 9 Blood and joint fluid are also acceptable specimen types for culture for detection of disseminated gonococcal infection.
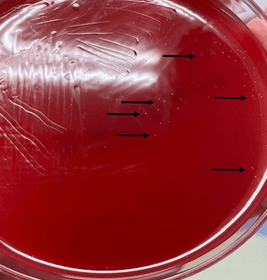
Enriched selective media for culture of N. gonorrhoeae includes MTM medium, ML medium, GC-Lect and the New York City medium. Plates should be incubated in a CO2 incubator (between 3-7%) at 35C to 37C for optimal growth.9 Gram negative diplococci recovered from urogenital sites that grow on the selective media and are oxidase-positive can be presumptively identified as N. gonorrhoeae. Another quick biochemical test that can be done is superoxol; N. gonorrhoeae produce immediate bubbling whereas N. meningitidis and N. lactamica produce weak, delayed bubbling. Confirmation using other testing methods such as carbohydrate utilization tests (e.g. N. gonorrhoeae produces acid from glucose only), immunological methods, enzymatic procedures, or DNA probe are also available.10
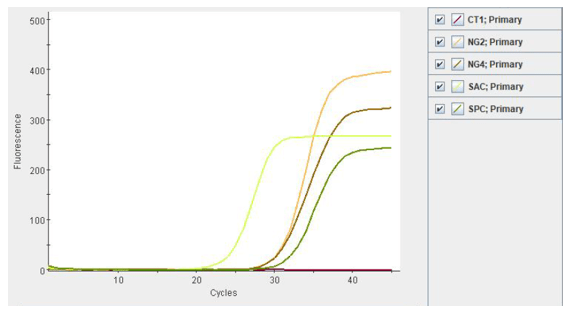
Compared to standard culture methods, Nucleic Acid Amplification Tests (NAAT) offer more rapid results and increased sensitivity. Additionally, NAATs may also include additional targets such as C. trachomatis, a frequent co-pathogen, as part of the assay. NAATs should be used according manufacturer’s protocols and on validated specimen types. For example, the Cepheid Xpert CT/NG test (as used by our patient here) can be used to test asymptomatic and symptomatic individuals and the acceptable specimen types are urine, pharyngeal, and rectal swabs, patient-collected vaginal swabs, and clinician-collected endocervical swabs.11 Given the legal implications of a N. gonorrhoeae diagnosis in a child, the CDC recommends that NAATs can be used to test for N. gonorrhoeae from vaginal and urine specimens from females and urine for males.12 For extragenital specimens, only validated FDA-cleared NAATs assays using pediatric specimens should be used.
The CDC recommends that uncomplicated gonorrhea be treated with ceftriaxone and azithromycin. However, between 2000-2010s, elevated MICs to both ceftriaxone and cefixime were seen and emerging azithromycin resistance is still a concern. The CLSI M100 currently recommends agar dilution or disk diffusion for antimicrobial susceptibility testing for N. gonorrhoeae. Susceptible and resistant interpretative breakpoints are available for penicillin, most cephems, tetracycline, ciprofloxacin, and spectinomycin. Of note, for azithromycin, only the susceptible category has a breakpoint.13

References
- CDC. Sexually Transmitted Disease Surveillance, 2020. Atlanta, GA: Department of Health and Human Services; April 2022.
- John J, Donald WH. Asymptomatic urethral gonorrhoea in men. Br J Vener Dis 1978; 54:322.
- Handsfield HH, Lipman TO, Harnisch JP, et al. Asymptomatic gonorrhea in men. Diagnosis, natural course, prevalence and significance. N Engl J Med 1974; 290:117.
- Sherrard J, Barlow D. Gonorrhoea in men: clinical and diagnostic aspects. Genitourin Med 1996; 72:422.
- McCormack WM, Johnson K, Stumacher RJ, Donner A, Rychwalski R. Clinical spectrum of gonococcal infection in women. Lancet, 1(8023), 1182–1185 (1977).
- Curran J, Rendtorff R, Chandler R, Wiser W, Robinson H. Female gonorrhea: its relation to abnormal uterine bleeding, urinary tract symptoms, and cervicitis. Obstet Gynecol, 45(2), 195–198 (1975).
- O’Brien JP, Goldenberg DL, Rice PA. Disseminated gonococcal infection: a prospective analysis of 49 patients and a review of pathophysiology and immune mechanisms. Medicine (Baltimore) 1983; 62:395.
- Laurer BA, Masters HB. Toxic effect of calcium alginate swabs on Neiserria gonorrhoeae. J Clin Microbiol 1988: 26:54-56
- Leber, A. 3.9 Genital Cultures. Clinical Microbiology Procedures Handbook, 4th Edition. ASM Press, Washington, DC. 2016. p.3.9.3.1-3.9.3.15.
- Knapp JS. Historical perspectives and identification of Neisseria and related species. Clin Microbiol Rev 1988;1:415-431.
- Cepheid GeneXpert. Xpert CT/NG (English). 2019. 301-0234 Rev.K
- CDC. Gonococcal Infections Among Infants and Children. Sexually Transmitted Infection Treatment Guidelines, Atlanta, GA: Department of Health and Human Services; 2021.
- CLSI. Performance Standards for Antimicrobial Susceptibility Test. CLSI supplement M100. Wayne, PA: Clinical and Laboratory Standards Institute; 2022, Edition 32

-Maikel Benitez Barzaga, MD is a Pathology Resident (PGY-1) at The George Washington University Hospital. His academic interest include hematology, microbiology, molecular and surgical pathology.

-Rebecca Yee, PhD, D(ABMM), M(ASCP)CM is the Chief of Microbiology, Director of Clinical Microbiology and Molecular Microbiology Laboratory at the George Washington University Hospital. Her interests include bacteriology, antimicrobial resistance, and development of infectious disease diagnostics.