A 4-year-old girl with no past medical history had been feeling unwell for one day following a barbecue she had attended a few days prior. Her symptoms worsened to include colicky abdominal pain and bloody diarrhea, with as many as eight bowel movements per day. This persisted for the following two days; thereafter, she presented to the hospital also complaining of fever, nausea, and vomiting. She was found to be dehydrated and pale on exam, and was admitted for intravenous rehydration. Fecal leukocyte testing and stool cultures were sent. A Gram stain of the pathogen isolated from stool culture is shown in Figure 1.
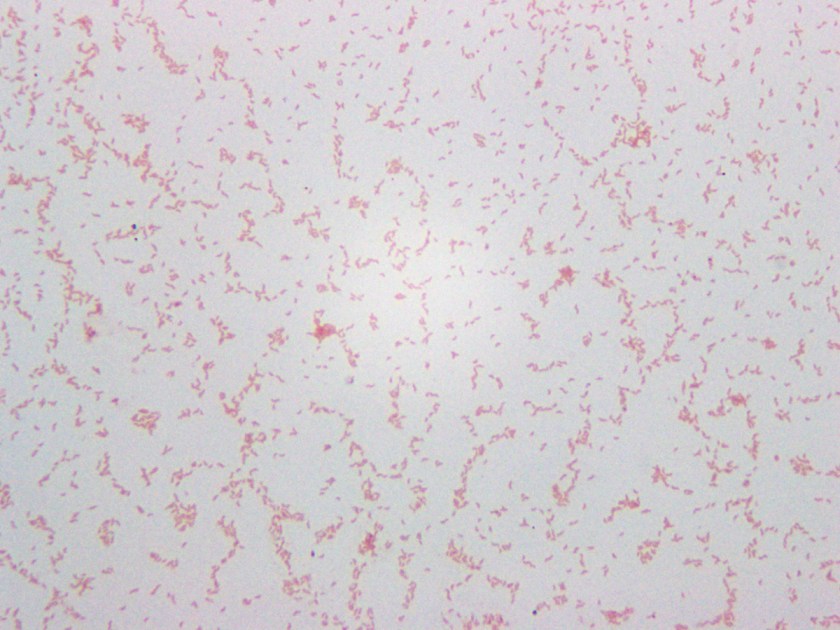
An infectious etiology is highly suspected given this patient’s presentation, leading to work-up with fecal leukocytes and stool cultures. The presence of fecal leukocytes, which was positive in this patient, is a strong indicator of inflammatory diarrhea. Bacterial stool culture allows for detection of Salmonella, Shigella, Campylobacter, E. coli O157:H7, Yersinia, Aeromonas, and Plesiomonas.
Many different culture mediums are used to isolate bacterial gastrointestinal pathogens. In addition to the routine 5% sheep blood agar and MacConkey agar, a case of infectious diarrhea requires further workup to rule out the above mentioned pathogens. Sorbitol-MacConkey agar is a variant of traditional MacConkey agar, and is used to detect E. Coli O157:H7, which differs from other E. coli strains by its inability to ferment sorbitol, thus forming colorless colonies on this media. Xylose lysine deoxycholate (XLD) and hektoen enteric (HE) agars are utilized for the selection and differentiation between Salmonella and Shigella. A sweep of bacteria growing on the blood agar plate and subsequent oxidase testing is used for detection of Aeromonas and Plesiomonas, which are oxidase positive organisms unlike normal fecal flora which is oxidase negative. Cefsulodin-irgasan-novobiocin (CIN) agar is used for the selection and differentiation of Yersinia, which utilizes inhibitory substances (cefsulodin, irgasan, novobiocin, bile salts, and crystal violet) to prevent the growth of most bacteria. The agar also contains a pH indicator that turns red or pink when mannitol is fermented; with Yersinia having a characteristic ‘bull’s eye’ colonies with red centers and clear edges. CIN is incubated at room temperature for 48 hours. Finally, Campy CVA agar is a selective media for Campylobacter containing antimicrobial agents cefoperazone, vancomycin, and amphotericin B (CVA) which inhibit normal fecal flora. This media is incubated at 42°C under microaerophilic conditions, which support the growth of Campylobacter jejuni and C. coli.
Our patient’s culture grew gray, non-hemolytic colonies on Campy CVA agar (Figure 2). The organism was identified as Campylobacter jejuni by MALDI-TOF MS (matrix-assisted laser desorption/ionization, time of flight mass spectrometry).

Campylobacter are gram-negative, microaerophilic, curved or spiral rods in the family Campylobacteriaceae. They are widely distributed in animals and infection is most often transmitted by contaminated foods, particularly undercooked chicken. The species most commonly associated with human infections are C. jejuni and C. coli, with C. jejuni accounting for the large majority. Infection with C. jejuni has been linked with subsequent development of Guillain-Barre syndrome two to three weeks following the initial illness. Our patient improved following two days of IV fluids and antibiotics with no subsequent follow up after discharge.
References:
Manual of Clinical Microbiology, 11th edition
-Said Albahra, MD, 1st year Anatomic and Clinical Pathology resident at the University of Texas Southwestern Medical Center.

-Erin McElvania TeKippe, Ph.D., D(ABMM), is the Director of Clinical Microbiology at Children’s Medical Center in Dallas Texas and an Assistant Professor of Pathology and Pediatrics at University of Texas Southwestern Medical Center.