Chromosome Structure
In a post back in September, I quickly summarized the abnormalities that can occur with chromosomes as a whole (such as deletions, insertions, transversions, etc). There is so much more to learn (more than I could possibly put into one blog post), because the way chromosomes behave, depends on their structure and DNA sequence. For instance, genes with the same DNA sequence will behave differently depending on where they are located on a chromosome as well as the effect of the surrounding DNA sequence.
So how exactly is the immense length of DNA compacted into a chromosome? Let’s take a DNA sequence and see just how it makes up a chromosome. A single molecule of DNA spools around histone protein cores forming bead like structures called nucleosomes. Between each nucleosome is a sequence of DNA termed “linker DNA.” The amino acids associated with histones are lysine and arginine. The super coiled form is compacted and can be visualized as a karyotype in laboratory testing.
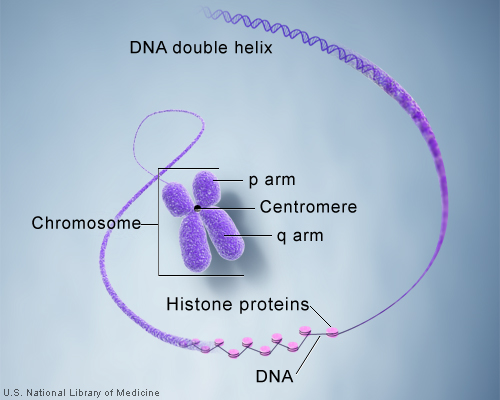
The centromere is the connection point of the duplicated chromosome, while telomeres are the endpoints. The short arm of the chromosome is termed “p” and the long arm of the chromosome is termed “q.” If we take these two chromosome arms into consideration, there are three types of chromosome morphology:
- Metacentric – Chromosome arms are equal in length
- Sub-centric – One arm is longer than the other
- Acro-centric / Telocentric – One arm is extremely small or even missing
Chromosomal Staining Methods
As I mentioned above the complete set of chromosomes for an individual can be visualized via a karyotype. I’ve listed a few of the ways this can be accomplished:
- G-Banding – Chromosomes are stained with giesma stains. The appearance differs based on the treatment of chromosomes prior to staining.
- Q-Banding – Chromosomes are stained with fluorescent dyes, quinacrine or quinacrine mustard. Q-Band staining is similar to G-banding in that the fluorescent regions represent the AT-rich regions of the chromosome.
- R-Banding – Results from heat treatment in a phosphate buffer followed by staining with Giesma dyes.
- C-Banding – Centromere staining that results from alkali treatment.
| TYPE OF BANDING | STAINING SUMMARY |
| G-Banding | · Geisma stain
· AT-rich regions stain darker than GC-rich regions |
| Q-Banding | · Quinacrine fluorescent dye stains AT-rich regions |
| R-Banding | · Banding pattern is opposite G-banding |
| C-Banding | · Stains heterochromatic regions close to the centromeres
· Usually stains the entire long arm of the Y chromosome |
So how do you exactly identify chromosomal location based on banding patterns?
In studying disease and mutation, we follow a specific type of nomenclature to designate the regions that are of interest to us. Let’s take for instance something like the 22q11.2 deletion. What do all of these numbers and letters mean? To quickly summarize, 22q11.2 deletion syndrome occurs from the deletion of a small piece of chromosome 22 at a location: q11.2
| 22q11.2 DELETION | |
| 22 | · Chromosome 22 |
| q | · Long arm of chromosome (q) |
| 1 | · Region 1 |
| 1 | · Band 1 |
| 2 | · Sub band 2 |
So now, when we add in karyotope information you might see something like the following:
46, XY, del(8)(q21)
When you break it down, it states the patient is male (XY) and has a deletion in the long arm (q) of chromosome 8 at region 2, band 1
Translocation nomenclature can get a little more confusing:
46, XX, t(3;12)(p12.1;p11)
This designates a female has a translocation between the short arms (p) of chromosomes 3 and 12 and region 1, band 2, sub band 1; and region 1 band 1 respectively.
An example of Down syndrome: 47, XX + 21 (Female has an extra chromosome 21)
An example of Klinefelter Syndrome: 47, XXY (Male with extra X chromosome)
What do “FISH” have to do with Molecular Biology?
FISH, an acronym for Fluorescent In-Situ Hybridization, is a method used to detect and visualize protein, RNA, and DNA structures in the cell. FISH analysis is a relatively fast method that provides great resolution as it incorporates fluorescent probes labeled for detection of specific regions, deletions, and translocations. The images below show the difference between FISH and Karyotype images.
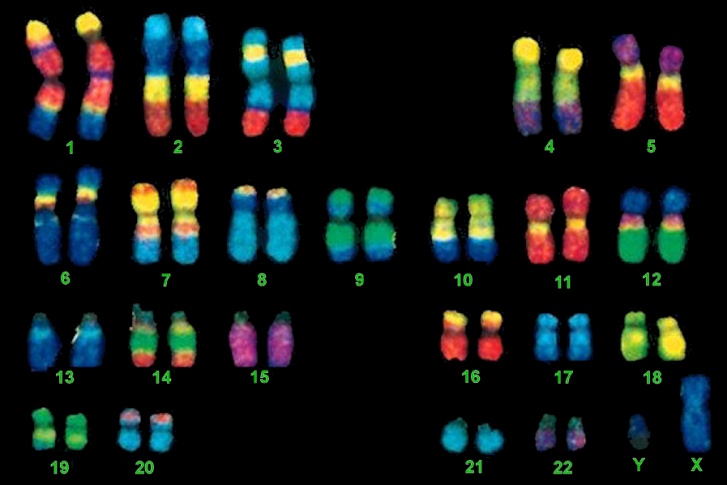
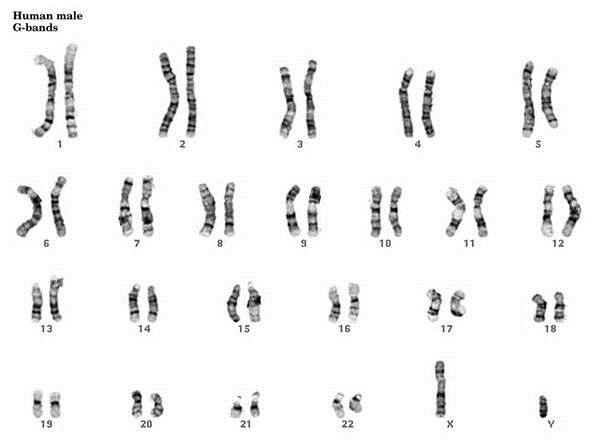
Robertsonian Translocations (ROB)
Robertsonian translocations are of importance because they involve translocating most of one entire chromosome to the centromere of another chromosome. They can be balanced or unbalanced. A balanced translocation usually results in no health difficulties because there isn’t a gain or loss of genetic material. However, due to the duplication or deletion of genetic material in an unbalanced translocation, syndromes and other malformations are likely to occur. The chromosome pairs common for Robertsonian translocations include translocations between 13 and 14, 14 and 21, and 14 and 15.
During a Robertsonian Translocation two chromosomes (typically acrocentric in formation) will break apart at their centromeres. The long arms will fuse to form a single chromosome and the short arms will also join to form a product. Typically the product created by the short arms contains nonessential genes and is eventually lost through cell division. Most people with ROB have only 45 chromosomes in each cell containing all of the essential genetic material and appear normal.
An example of a balanced Robertsonian Translocation would be when the long arms of chromosomes 14 and 21 fuse together. Phenotypically, the heterozygous carrier would appear normal because there are two copies of the major chromosome arms, resulting in two copies of the essential genes. However, children of the carrier could inherit an unbalanced translocation that causes Trisomy 21 (Down Syndrome).
Test your Knowledge!
- Which two amino acids are associated with histones?
- What type of chromosome morphology is shown:
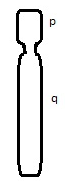
- If you wanted to stain chromosomes to see varying regions that were AT-rich, which type of stain would you use?
- Describe the following karyotype results: 46, XX, t(1;14)(p21.3; p17.6)
Answers
- Lysine and Arginine
- Sub-centric
- G-Banding (although Q-Banding also will produce darker regions that are AT-rich)
- Female patient, with a translocation between the p and q arms of chromosomes 1 and 14, and region 2, band 1, sub band 3; and region 1 band 7, sub band 6 respectively.
References:
Buckingham, L. (2012). Molecular Diagnostics: Fundamentals, Methods and Clinical Applications (2nd ed.). Philadelphia: F.A. Davis Company.
Coleman, W.B, Tsonagalis, G.J. (2005). Molecular Diagnostics: For the Clinical Laboratorian. New York: Springer-Verlang
Searle, B. Rarechromo.org. The Rare Chromosome Disorder Support Group. 1996. Web. 19 Dec. 2015.

-LeAnne Noll, BS, MB(ASCP)CM is a molecular technologist at Children’s Hospital of Wisconsin and was recognized as one of ASCP’s Top Five from the 40 Under Forty Program in 2015.

It’s very interesting subjects. Thank you, madam.
It is great information. thanks for posting it.