
Ah, the blue chip – not much fun to see after spending a day preparing the libraries and running clonal amp overnight. There are a couple possible explanations for a blue chip, and you can figure them out by looking at the metrics of the run.
Test Fragments
The test fragments serve as a control for the sequencing run. They are spiked into the mixture of library ISPs before they are loaded on the chip. These will allow you to figure out where the problem occurred if you encounter a blue chip. If the Test Fragments are detected and are of sufficient quality, then this means the sequencing run worked and the problem most likely occurred before sequencing, during library prep or clonal amplification. If the Test Fragments are not detected, then it could mean one of two things – one – the clonal amplification did not work for either the library or the Test Fragment ISPs, or – two – the sequencing run was somehow at fault. Let’s take a look at both examples.
Troubleshooting a Blue Chip
In the event you see a blue chip, first, check to see what kinds of ISPs showed up after the analysis. For the chip pictured above, there were ISPs that had product on them, as you can see in the Live category (6,475,553 ISPs or 95.3% of the ISPs, shown in the screenshot below). This means clonal amp was successful for a small number of the library ISPs. Next, there were also Test Fragments detected, at 433,392 ISPs or 6.7% of the total ISPs. Scroll down to the bottom of the page, and you will see how the Test Fragments sequenced. We like to see the Percent 50AQ17 and Percent 100AQ17 at least in the 80’s, but even still, you can see that these were detected and were sequenced. Because of this, the sequencing run looks to be fine, so most likely the problem occurred before sequencing. In this case, we believe the library prep did not yield the expected 100pM concentration, so the library pool was over-diluted prior to clonal amplification. The library prep was repeated, and clonal amplification was run on the new pool of libraries, and the sequencing was successful.


In this next example, we have the other possibility. This chip was blue as well (this is a 520 chip, instead of a 530, to explain the different sized pictures).
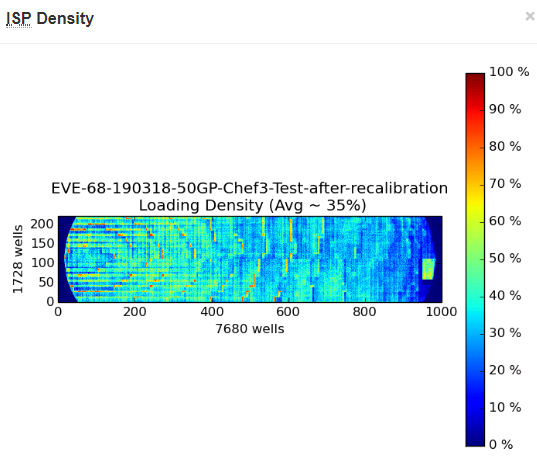
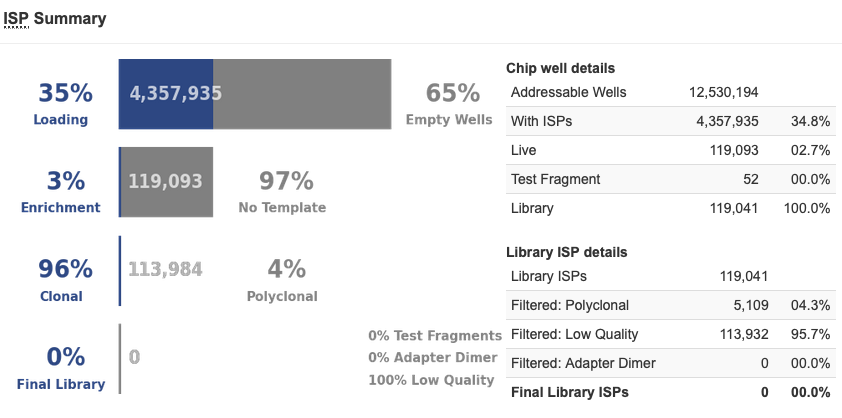
First, there are only 2.7% Live ISPs, so even lower than the chip above. But the even stranger thing was that there were 0.0% Test Fragments, and at the end of the analysis, there were absolutely no ISPs left to be analyzed, library or Test Fragment. This was the only time we had ever seen a chip like this; generally, if we had blue chips, they were like the previous example. We looked at our library pool quant and it was in the expected range, so we did not believe it was a library prep issue. The sequencing initialization was successful and did not have any errors, so we did not believe it was a sequencing problem. We repeated clonal amplification with the same library pool and had successful sequencing. In speaking with our Field Application Scientist, it was decided it must have been a failure of one of the reagents of the clonal amplification – either a Taq was not present or something, so the clonal amplification never occurred, or something similar.
Hopefully you will not experience too many of these blue chips, but if you do, I hope you are a little more prepared to troubleshoot! Happy sequencing!

-Sharleen Rapp, BS, MB (ASCP)CM is a Molecular Diagnostics Coordinator in the Molecular Diagnostics Laboratory at Nebraska Medicine.

