Case History
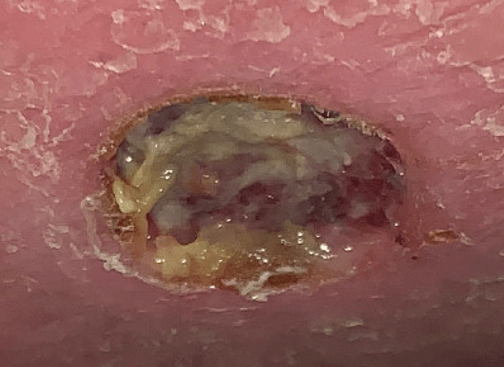
A middle-aged man with osteogenesis imperfecta, poorly controlled HIV, and hepatitis C presented for outpatient management of an infected non-pressure ulcer on his ankle. The patient had no history of recent travel and was current on vaccinations. He was afebrile but reported increasing difficulty with ambulation due to pain from the lesion. Per the patient, the wound began as an itchy “bug bite” two weeks prior, which he had scratched, causing skin breakage. The ulcer became progressively larger with corresponding increases in pain, warmth and swelling. Specimens from the ulcer were collected for culture, and the patient started empiric doxycycline in addition to prophylactic amoxicillin/clavulanate he was already taking for management of a pre-existing ulcer on the opposite foot. The non-healing wound progressed to the size of a nickel with worsening pain. Due to these symptoms and the associated microbiological data, the patient was instructed to present to the emergency department where he was admitted for additional evaluation. Upon admission, the ulcer exhibited no surrounding erythema, but a slight exudate and pitting edema was noted (Image 1). An X-ray of the ankle was obtained which revealed soft tissue swelling, but no fracture or crepitus.
Microbiology
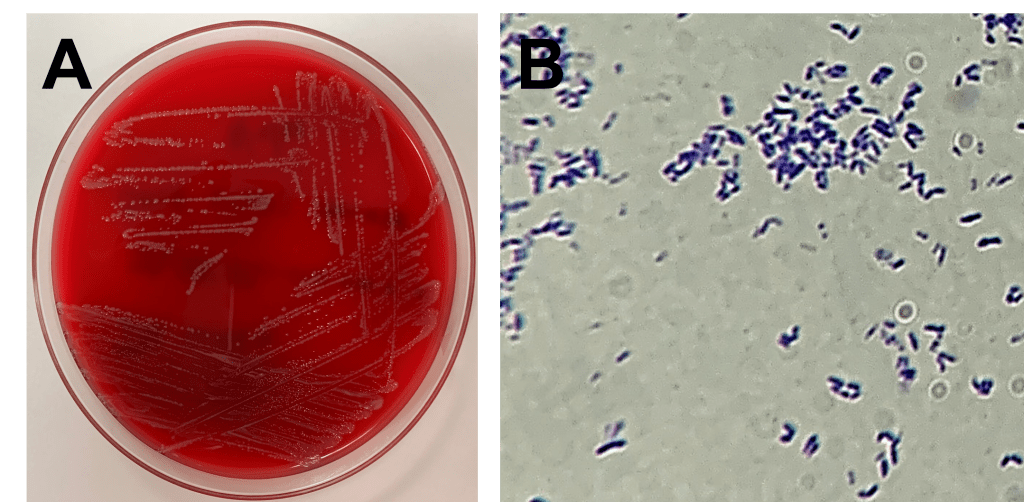
Specimens of the ulcerated lesion were submitted to the microbiology laboratory for routine bacterial culture. No growth was observed on MacConkey agar plates, while two beta-hemolytic morphotypes and one non-hemolytic morphotype were observed on blood agar. The two beta-hemolytic species were identified as Streptococcus dysgalactiae and Arcanobacterium haemolyticum by MALDI-TOF MS. The third, non-hemolytic organism was a catalase-positive, gram positive coryneform rod (Image 2A and B), and was identified by MALDI-TOF MS as Corynebacterium diphtheriae. This identification was subsequently confirmed by both a commercial reference laboratory and the US Centers for Disease Control and Prevention.
Discussion
Corynebacterium diphtheriae is the etiological agent of diphtheria, a toxin-mediated disease classically associated with respiratory and cutaneous infections. C. diphtheriae is infrequently encountered in the United States due to a robust national vaccination program but remains endemic in other parts of the world. Respiratory manifestations include pharyngitis with dysphagia, lymphadenitis (often described as a “bull-neck” appearance), and the development of a characteristic white/grey pharyngeal pseudomembrane which can cause airway obstruction. The pathogenesis of C. diphtheriae is mediated by diphtheria toxin which inhibits host cell protein synthesis leading to cell death. Diphtheria toxin can also cause cardiac, nephrotic, and neurological sequalae due to dissemination. The diphtheria toxin gene (tox) is encoded on a bacteriophage which lysogenizes into the bacterial chromosome and is expressed in response to low iron concentrations. While C. diphtheriae is most frequently associated with diphtheria toxin production, Corynebacterium ulcerans and Corynebacterium pseudotuberculosis can also express the toxin if infected with the bacteriophage.
Cutaneous C. diphtheriae infections manifest initially as a vesicle, eventually developing into a painful ulcerative lesion that may or may not have a pseudomembrane. These infections can be caused by either fully toxigenic strains, non-toxigenic strains lacking the tox gene, or non-toxigenic toxin gene bearing (NTTB) strains.1 NTTB strains are genotypically positive for the presence of the tox gene on the lysogenized phage, but do not express functional diphtheria toxin. This can be due to 1) mutation of truncation of the tox gene coding sequence, 2) promotor mutations, or 3) alterations in proteins regulating tox expression. NTTB strains are important from an epidemiological perspective in that they serve as an environmental reservoir for tox gene-harboring phage which could convert circulating non-toxigenic C. diphtheriae into toxin-producing organisms.1-3 This phenotype also presents additional diagnostic challenges as toxin gene expression must therefore be confirmed by more laborious phenotypic methods instead of genotypically (i.e. by PCR).
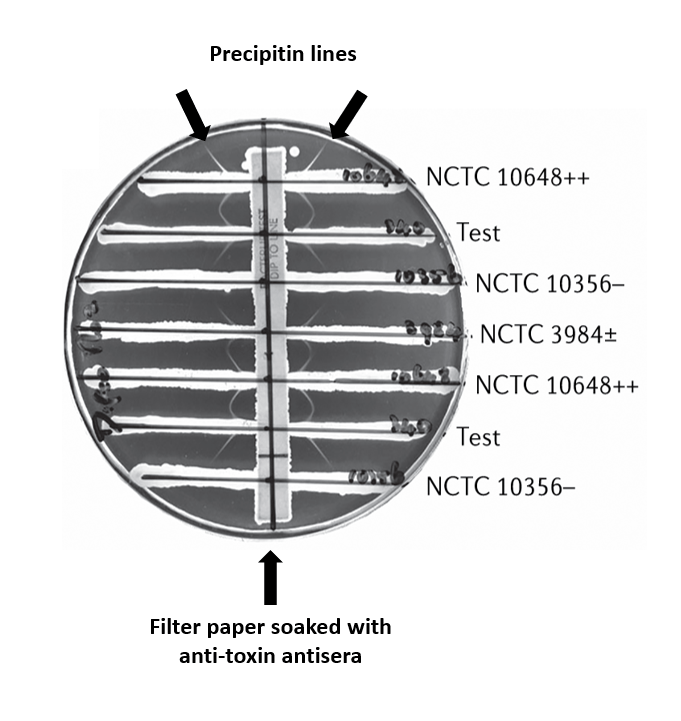
Recovery of C. diphtheriae in the routine setting is challenging as the organism morphologically resembles other coryneform rods usually representative of flora in cutaneous and respiratory specimens. While selective and differential medias are available (i.e. Cystine Tellurite Blood Agar or Tinsdale medium), they are not used routinely due to low incidence. As both toxigenic and non-toxigenic strains of C. diphtheriae circulate, this isolate was referred to the CDC’s Pertussis and Diphtheria laboratory for additional typing and toxin analysis. The organism was determined to belong to the mitis biotype and was positive for the tox gene by PCR. Toxin gene expression was then evaluated by the Elek Immunodiffusion test (Image 3). In this classical method, a filter paper strip saturated with anti-toxin is placed perpendicular to control and test strains of the organism on non-selective media. If the organism expresses diphtheria toxin, the toxin and the antisera form a complex and precipitate from solution. This phenomenon is visualized as precipitin lines in the agar after 24 hours of incubation at 37°C.4 Despite tox gene PCR-positivity, the Elek Immunodiffusion test revealed that this patient’s isolate did not express diphtheria toxin. Thus, this C. diphtheriae isolate was a representative example of a NTTB strain.
In contrast to respiratory presentations which have declined due to vaccination, cutaneous infections with C. diphtheriae have become more frequently recognized. Unlike respiratory diphtheria, cutaneous diphtheria was not reportable to the National Notifiable Diseases Surveillance System until the clinical criteria changed in 2019. Since that time, toxigenic isolates recovered from either respiratory or cutaneous sources are reportable.5 Additionally, the incorporation of MALDI-TOF MS into routine workflow has facilitated identification of the organism when isolated clinically and likely increased reporting.
Although immunization protects against clinical diphtheria, it does not prevent colonization by non-toxigenic C. diphtheriae including NTTB strains.2,3 Non-toxigenic C. diphtheriae causing cutaneous infections are often recovered with other pyogenic organisms including Staphylococcus aureus, beta-hemolytic streptococci, and A. haemolyticum6 as was observed in this case. Risk factors for cutaneous infections include a compromised immune system, eczema, travel to endemic regions, intravenous drug use, homelessness/unsanitary living conditions, and alcoholism.3,7 It is unclear how this patient was exposed, although he did have risk factors including immunosuppression and those associated with his osteogenesis imperfecta. The patient was started on erythromycin and ampicillin/sulbactam for subsequent management and has continued to improve when seen at follow-up. Subsequent cultures of the wound have remained negative for C. diphtheriae.
- Zakikhany K, Neal S, Efstratiou A. 2014. Emergence and molecular characterisation of non-toxigenic tox gene-bearing Corynebacterium diphtheriae biovar mitis in the United Kingdom, 2003–2012. Eurosurveillance 19:20819.
- Doyle CJ, Mazins A, Graham RMA, Fang N-X, Smith HV, Jennison AV. 2017. Sequence Analysis of Toxin Gene-Bearing Corynebacterium diphtheriae Strains, Australia. Emerging infectious diseases 23:105-107.
- Sharma NC, Efstratiou A, Mokrousov I, Mutreja A, Das B, Ramamurthy T. 2019. Diphtheria. Nature Reviews Disease Primers 5:81.
- Kates O, Starr K, Bourassa L, Burnham C-AD. 2020. The Brief Case: Nontoxigenic Corynebacterium diphtheriae in a Nonhealing Wound. Journal of Clinical Microbiology 58:e00506-00520.
- United States Centers for Disease Control and Prevention. 2019. Diphtheria (Corynebacterium diphtheriae) 2019 Case Definition – National Notifiable Diseases Surveillance System. https://ndc.services.cdc.gov/case-definitions/diphtheria-2019/. Accessed August 1st, 2021.
- Lowe CF, Bernard KA, Romney MG. 2011. Cutaneous Diphtheria in the Urban Poor Population of Vancouver, British Columbia, Canada: a 10-Year Review. Journal of Clinical Microbiology 49:2664-2666.
- Gubler J, Huber-Schneider C, Gruner E, Altwegg M. 1998. An Outbreak of Nontoxigenic Corynebacterium diphtheriae Infection: Single Bacterial Clone Causing Invasive Infection Among Swiss Drug Users. Clinical Infectious Diseases 27:1295-1298.

-Andrew Clark, PhD, D(ABMM) is an Assistant Professor at UT Southwestern Medical Center in the Department of Pathology, and Associate Director of the Clements University Hospital microbiology laboratory. He completed a CPEP-accredited postdoctoral fellowship in Medical and Public Health Microbiology at National Institutes of Health, and is interested in antimicrobial susceptibility and anaerobe pathophysiology.
