Hello everyone, it’s been quite some time since my last post. I hope everyone has remained safe and healthy during these times!
My last post dived into short tandem repeat (STR) analysis for bone marrow engraftment monitoring. Today is a presentation of a patient who was transplanted for treatment of acute myeloid leukemia (AML). With all patients (with minor exceptions), donor and pre-transplant recipient samples are taken before transplant. Their informative alleles are then identified and used to determine the percent of donor and any recipient cells in subsequent post-transplant samples.
This patient was unique in that we were not able to obtain the donor sample (they were transplanted outside of our system), and therefore we used a buccal swab for their pre-transplant recipient informatives.
Buccal swabs are chosen because they are a non-invasive way to obtain squamous epithelial cells. These cells are important because they are of the recipient origin and will not change. With this technique, it is essential that the patient has no mucosal inflammation or is not too rough when swabbing their cheek. Otherwise, the buccal sample may become contaminated with blood which would contain donor cells.
We then inferred the donor informatives from the data of a mixed sample and the buccal swab.
Calculation of recipient and donor percentage in a post-transplant sample is determined on specific formulas that utilize these informative alleles. But what happens when a patient relapses and new mutations or deletions are introduced into their genome, causing a change in these informative alleles?
In this case, the patient had a loss of allele at two loci (CSF1PO – allele 11 and D5S818 – allele 13) after having previously obtained full engraftment (Figure 1).
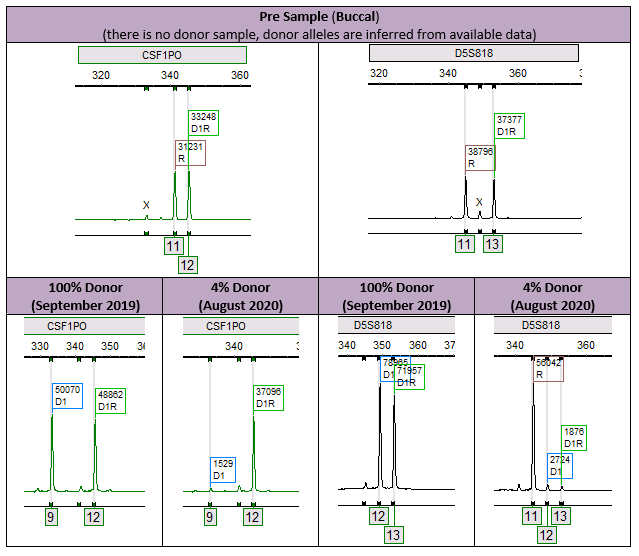
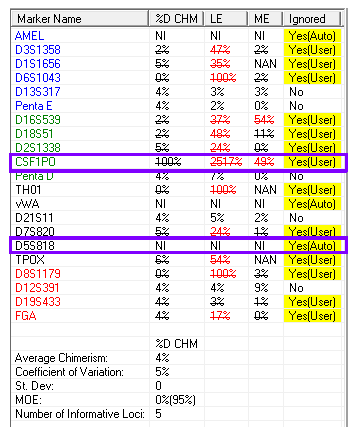
The importance here is that the true percent donor is 4% (Figure 2). If we take a look at the affected informative alleles, we see an erroneous result of 100% donor and NI (which means the locus is non-informative, eliminating it from the calculations). This expands on the importance of an analyst to be attentive to the results presented. While this case was clearly evident and was caught by our error measurements, it is theoretically possible to cause an issue, especially in cases where the recipient percentages may be smaller. Furthermore, this phenomenon stresses the importance of including multiple informative alleles in our analysis, which increases our measurement of confidence.1
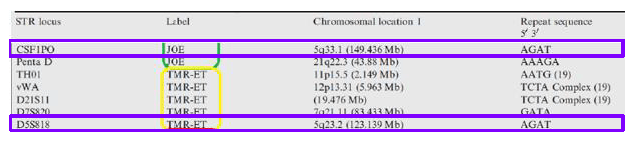
We know that a loss of allele (loss of heterozygosity) is the likely explanation because both loci are in locations specific to the disease. Looking at Figure 3 below, the two alleles were affected because they were both present on the long arm of chromosome 5. Further, this chromosome is known to be involved in AML, and is also, of course, associated with other disorders like MDS.2 Additionally, the patient had cytology testing that identified this as an affected chromosome.
This is an interesting phenomenon and one that shows in measurable terms how a patient’s status can affect their molecular results. It’s further an expression of the molecular mechanisms of a disease, one of my first measurable experiences of how a disease affects the physical molecular constituents of another human.
To me, this encounter was an expression of how complicated, and yet connected, the entire genome has been designed. I am continuously amazed and look forward to expanding my understanding of molecular science.
References
- Crow J, Youens K, Michalowski S, et al. Donor cell leukemia in umbilical cord blood transplant patients: a case study and literature review highlighting the importance of molecular engraftment analysis. J Mol Diagn. 2010;12(4):530-537. doi:10.2353/jmoldx.2010.090215
- Crow J, Youens K, Michalowski S, et al. Donor cell leukemia in umbilical cord blood transplant patients: a case study and literature review highlighting the importance of molecular engraftment analysis. J Mol Diagn. 2010;12(4):530-537. doi:10.2353/jmoldx.2010.090215

-Ben Dahlstrom is a recent graduate of the NorthShore University HealthSystem MLS program. He currently works as a molecular technologist for Northwestern University in their transplant lab, performing HLA typing on bone marrow and solid organ transplants. His interests include microbiology, molecular, immunology, and blood bank.
