Case Report
A 77 year old male presented to the hospital with chest pain, lightheadedness, burning urination for the past few weeks. He has blood in his urine due to a previously diagnosed neoplasm. The patient moved from India to the United States in February, with a diagnosis of bladder cancer and a history of hypertension, congestive heart failure, coronary artery disease, and atrial fibrillation. In the hospital, abscesses on both right and left kidneys were found, and patient had nephrostomy tubes placed. Purulent discharge confirmed he had a severe urinary tract infection.
Laboratory Identification
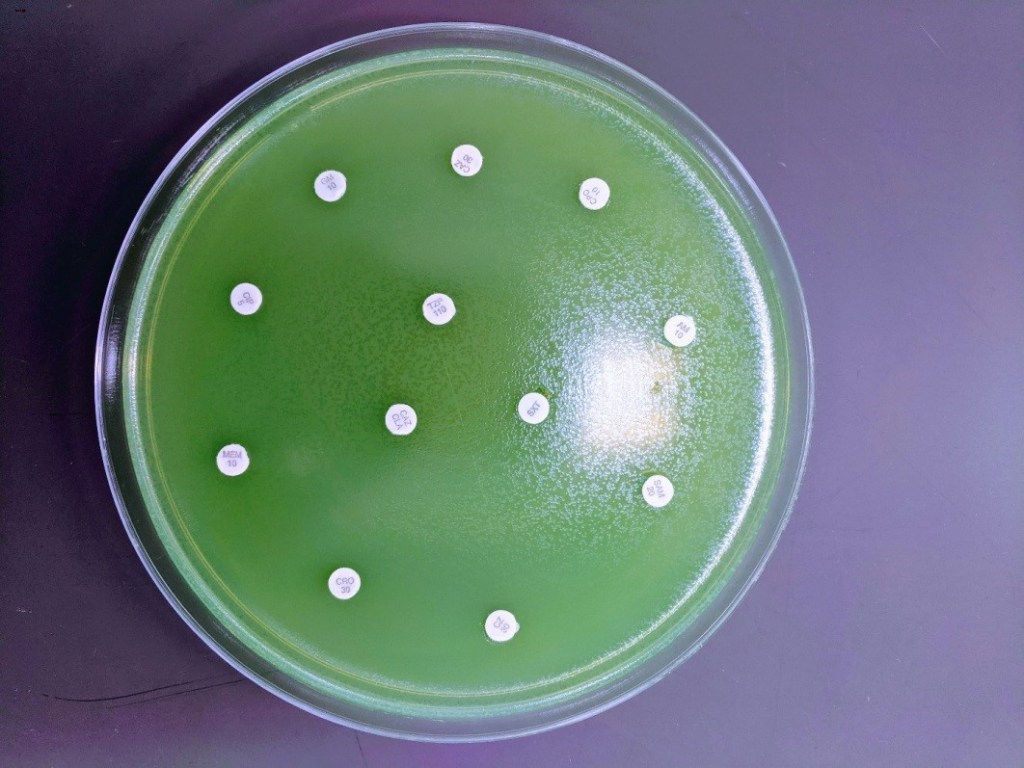
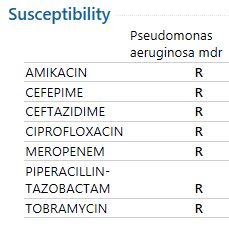
The patient’s urine culture grew >100,000 colony forming units/milliliter (CFU/ml) of an oxidase-positive, non-lactose fermenting Gram-negative rod. On the blood agar plate, large gray, smooth, flat, mucoid, β-hemolytic colonies were found. Although bacteria growing on solid media should not be actively smelled, the organism emitted a grape or tortilla smell from the plate. The organism was identified as Pseudomonas aeruginosa by MALDI-TOF mass spectrometry. The isolate was plated onto Mueller Hinton agar for Kirby-Bauer disc diffusion antibiotic susceptibility testing (Image 1). A fluorescent green lawn of bacteria grew up to the edge of all discs, indicating high-level resistance to all antibiotics tested (Table 1). Modified carbapenem inactivation method (mCIM) testing was positive and Cepheid GeneXpert CarbaR PCR testing revealed that this P. aeruginosa isolate carried the New Delhi metallo-β-lactamase-1 carbapenemase (NDM-1).
Discussion
The issue of super bugs is on the rise, with the fear of antibiotic resistance disseminating through more bacterial populations and species. Carbapenems are drugs that are very powerful broad-spectrum antibiotics, usually reserved as a last resort treatment for serious and resistant infections.1 β-lactamases are divided into four Ambler classes: A, B, C, and D. Class B differs from the others because it utilizes zinc as a metal cofactor for its catalytic activity. The others use a serine residue for their catalytic activity.2
NDM-1 is a class B β-lactamase. It was named after New Delhi, India when a Swedish resident presented with an extremely resistant infection after a trip to India in 2008. NDM-1 bacteria can now be found with high prevalence in India and China, and increasingly in other countries such as the UK and US.3,4 While the origination of the gene may not have been India, many of these infections are from people who have traveled to India or other Asian continents.5 Concerns about overprescribing and misuse of antibiotics in India are rising, where India is one of the biggest consumers of antibiotics in the world. One study even found striking evidence of this misuse, demonstrating that 2 out of 3 adults under 20 presented antibiotic resistance isolates to fluoroquinolones and/or cephalosporins.6,7,8
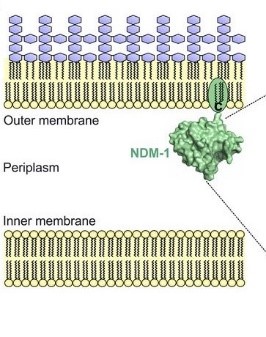
The gene for NDM-1 is blaNDM-1 and has been found on both plasmid and chromosomal components of different bacteria. Due to its presence on plasmids, the gene can easily spread through bacterial populations and other bacterial species – as has already been documented in Enterobacteriaceae and A. baumannii.3 The β-lactamase that it codes for is a lipoprotein that is anchored in the outer membrane of the gram negative bacteria. Other metalo-β-lactamases (MBLs) are periplasmic proteins, which are more affected by changes in essential metal cofactors in their enzymatic function. Thus far, it has been found that there are 16 discovered variants of NDM. Some variants being more fit than NDM-1. It is hypothesized that these variants are being selected for in the clinical setting, with the protein being more stable and demonstrating higher affinities for zinc during time of metal-chelating (a process the immune system adapts to combat infections).9 Unfortunately, NDM-1 and its variants are resistant to almost all antibiotics. Usually the only option is colistin and tigecycline.3
The disturbing issue, and the big picture, is the capability of MDR organisms and their genes of disseminating. As previously mentioned, NDM-1 is capable of spreading to other species and within its population. Yet, a terrifying report has demonstrated blaNDM-1 detection in artic soil samples from 2013, 4 years after the first detection of the gene.10 This demonstrates the ability for antibiotic resistance to spread on a global scale, and how serious this battle truly is.
References
- “Carbapenem-Resistant Enterobacteriaceae (CRE) Infection.” Centers for Disease Control and Prevention, Centers for Disease Control and Prevention, 23 Feb. 2015, http://www.cdc.gov/hai/organisms/cre/cre-patientfaq.html.
- Walther-Rasmussen, Jan, and Niels Høiby. “Class A Carbapenemases.” Journal of Antimicrobial Chemotherapy, vol. 60, no. 3, 2007, pp. 470–482., doi:10.1093/jac/dkm226.
- Khan, Asad U., et al. “Structure, Genetics and Worldwide Spread of New Delhi Metallo-β-Lactamase (NDM): a Threat to Public Health.” BMC Microbiology, vol. 17, no. 1, 2017, doi:10.1186/s12866-017-1012-8.
- Mohapatra P. R. (2013). Metallo-β-lactamase 1–why blame New Delhi & India?. The Indian journal of medical research, 137(1), 213–215.
- Roos, Robert. “Canada Finds More Infections with NDM-1 Resistance Factor.” University of Minnesota, Center for Infectious Disease Research and Policy, 11 Nov. 2010, http://www.cidrap.umn.edu/news-perspective/2010/11/canada-finds-more-infections-ndm-1-resistance-factor.
- Gupta, M., Didwal, G., Bansal, S., Kaushal, K., Batra, N., Gautam, V., & Ray, P. (2019). Antibiotic-resistant Enterobacteriaceae in healthy gut flora: A report from north Indian semiurban community. The Indian journal of medical research, 149(2), 276–280. doi:10.4103/ijmr.IJMR_207_18
- Kotwani, Anita, and Kathleen Holloway. “Access to Antibiotics in New Delhi, India: Implications for Antibiotic Policy.” Journal of Pharmaceutical Policy and Practice, vol. 6, no. 1, 2013, doi:10.1186/2052-3211-6-6.
- Kotwani, Anita, et al. “Antibiotic-Prescribing Practices of Primary Care Prescribers for Acute Diarrhea in New Delhi, India.” Value in Health, vol. 15, no. 1, 2012, doi:10.1016/j.jval.2011.11.008.
- Bahr, Guillermo, et al. “Clinical Evolution of New Delhi Metallo-β-Lactamase (NDM) Optimizes Resistance under Zn(II) Deprivation.” Antimicrobial Agents and Chemotherapy, vol. 62, no. 1, 2017, doi:10.1128/aac.01849-17.
- Mccann, Clare M., et al. “Understanding Drivers of Antibiotic Resistance Genes in High Arctic Soil Ecosystems.” Environment International, vol. 125, 2019, pp. 497–504., doi:10.1016/j.envint.2019.01.034.

-Ben Dahlstrom is a recent graduate of the NorthShore University HealthSystem MLS program. He currently works as a molecular technologist for Northwestern University in their transplant lab, performing HLA typing on bone marrow and solid organ transplants. He graduated with a bachelors in Biology at the University of Illinois at Chicago (UIC) and concurrently from the UIC Honors College. He discovered his passion for the lab through his experience in healthcare. His interests include microbiology, molecular, immunology, and blood bank.

-Erin McElvania, PhD, D(ABMM), is the Director of Clinical Microbiology NorthShore University Health System in Evanston, Illinois. Follow Dr. McElvania on twitter @E-McElvania.
