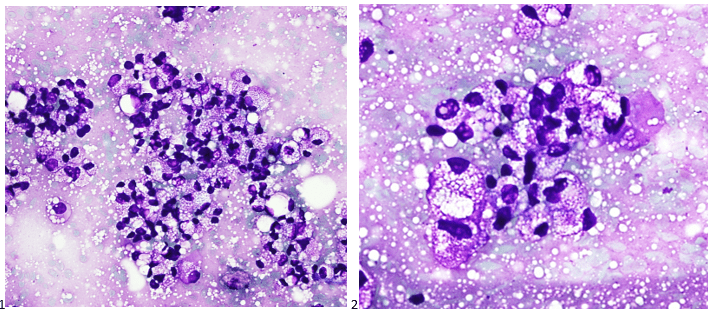
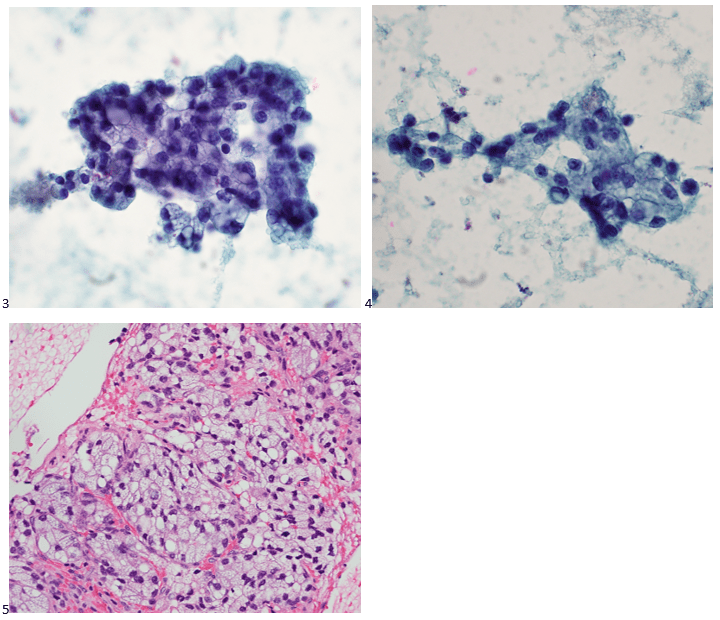
Case History
A 70 year old female with a medical history of treatment-related acute myeloid leukemia with absolute neutropenia due to chemotherapy presented to our emergency department with a two-day history of fever (Tmax 102°C) with progressively increasing erythema and tenderness reported in her right elbow around a healing surgical incision. Approximately one month prior to presentation the patient underwent surgical excision of her right upper extremity (RUE) cephalic vein for supportive thrombophlebitis as a complication of peripheral IV insertion. Additionally, two weeks prior to presentation the patient was noted to have three small areas (< 1 cm) of wound dehiscence along the surgical incision without overt signs of infection at her outpatient surgical follow-up visit (Figure 1A). These wounds were packed with iodoform gauze and wrapped with sterile kerlix gauze wrap followed by an elastic wrap. The patient was instructed to change the packing and dressing one to two times daily; however, the patient reported non-compliance with these instructions and the packing and dressing had not been changes in the two weeks from the clinic visit until ED presentation.

The patient was admitted to the hospital. Blood cultures were drawn and IV vancomycin, cefepime and metronidazole were initiated for empirical antimicrobial therapy in the setting of neutropenic fever and RUE cellulitis. 18 hours after collection both sets of blood cultures grew a gram positive rod. The infection diseases service was subsequently consulted to determine the significance of the gram positive rods growing in blood culture, if it represented contaminant vs a true pathogen. Imaging of the arm was recommended to further evaluate the extent of the RUE skin & soft tissue infection. CT imaging of the humerus and forearm revealed a thick-walled fluid collection concerning for abscess formation along the operative site involving both the arm and forearm of the patient (Fig. 2). The plastic surgery service was also consulted and noted old iodoform gauze in the areas of wound dehiscence with surrounding erythema (Fig. 1B); however, no purulence was noted from these areas. The erythema surrounding the wound continued to progress (Fig. 1C) during the hospitalization and the patient remained febrile despite the broad-spectrum antibiotic regiment. Given the patient treatment failure by medical management, the patient was taken for surgical washout and debridement of the wound. Deep cultures were obtained during the surgery. Both the tissue culture and blood cultures would grow Corynebacterium jeikeium. The patient developed a maculopapular cutaneous rash several days into the admission which was deemed to be an antibiotic-induced rash. The antimicrobial regiment was narrowed down to daptomycin due to the concern that vancomycin and/or cefepime may be responsible for the drug rash. The cellulitis and abscess resolved following surgical debridement and treatment with vancomycin/daptomycin; however, the patient elected to pursue comfort care status due to her underlying malignancy. She was transitioned to inpatient hospice where she would die shortly afterwards due to progression of her AML.

Laboratory
Corynebacterium species are non-motile, facultatively anaerobic, club-shaped gram positive rods that have a characteristic picket fence appearance when stained due to snapping division. These organisms are commonly encountered in clinical samples; however, given their low virulence in general and prevalence as common skin flora they are often considered contaminants or colonizers. The clinical significance of diphtheria toxin (DT)-producing strains has been known for decades. Non-DT-producing Corynebacterium species were historically considered as non-virulent but in recent years several non-DT-producing Corynebacterium species have been noted to be pathogenic. The wide spread availability of MALDI-TOF MS in most clinical microbiology labs has allowed for rapid and accurate species level identification of Corynebacterium allowing for identification of these pathogenic species in clinical settings. Some Corynebacterium species are highly lipophilic due to the presence of fatty acids, such as mycolic acid, in their cell wall. Isolation of lipophilic Corynebacterium species on standard culture media can be challenging but can be accomplished utilizing blood agar, due to the presence of lipid-containing red cell membranes, with at least 48 hours of incubation. For optimal recovery of lipophilic strains agars supplemented with Tween 80 produce the best recovery rates.
As a lipophilic species, C. jeikeium can be difficult to identify and propagate. It does not grow well on chocolate agar (Fig. 3A & 3C). On blood agar it tends to have scant or dusky growth at 24 hours (Fig. 3B) and appear as translucent, pinpoint colonies at 48 hours (Fig. 3D). Identification using MALDI-TOF can be challenging before 48 hours due to its poor growth on standard media. For comparison, C. striatum, a non-lipophilic Corynebacterium spp. grows relatively well on both blood and chocolate agar. Several Corynebacterium species, including C. jeikeium and C. striatum, are known to be multidrug resistant, especially towards beta-lactam antibiotics.

Discussion
Corynebacterium jeikeium was initially identified in 1976 and classified by CDC as group JK diptheroids. It is considered part of normal skin flora like most other Corynebacterium spp. In immunosuppressed individuals, especially in persons with hematolymphoid malignancies, invasive disease including dissemination infections have been reported. Infections can also be seen in hardware-associated and prosthetic joint-related infections. Due to its predilection for biofilm formation, C. jeikeium can be difficult to treat in these cases. Cases of infective endocarditis have been reported as well. Like many Corynebacterium species, C. jeikeium is often multidrug resistant. Vancomycin is the first line agent for treatment of infections due to C. jeikeium. Linezolid and daptomycin are other alternative treatment options. When isolating C. jeikeium from blood sources especially in immunocompromised patients like in our case, physicians and other health-care providers must thoroughly assess the total clinical picture before dismissing it as a contaminant.
References
1. Bernard K., Identification of Gram-Positive Bacteria (2023) in AL Leber & CAB Burnham (Eds.) Clinical Microbiology Procedures Handbook (5th ed.) doi:10.1128/9781683670438.CMPH.ch3.18
2. Gupta R, Popli T, Ranchal P, Khosla J, Aronow WS, Frishman WH, El Khoury MY. Corynebacterium Jeikeium Endocarditis: A Review of the Literature. Cardiol Rev. 2021 Sep-Oct 01;29(5):259-262. doi: 10.1097/CRD.0000000000000355. PMID: 32976125.
3. Mattos-Guaraldi AL, Sanchez Dos Santos L, & Vieira VV, Coryneform Gram-Positive Rods (2023) in Carroll et al. (Eds.) Manual of Clinical Microbiology (15th ed.) doi:10.1128/9781683670438.MCM.ch28
4. Moore Pardo SM, Patel RH, Ramsakal A, Greene J. Disseminated Corynebacterium jeikeium Infection in Cancer Patients. Cureus. 2020 Jun 22;12(6):e8764. doi: 10.7759/cureus.8764. PMID: 32714702; PMCID: PMC7377673.
5. Murray BE, Karchmer AW, Moellering RC Jr. Diphtheroid prosthetic valve endocarditis. A study of clinical features and infecting organisms. Am J Med. 1980 Dec;69(6):838-48. doi: 10.1016/s0002-9343(80)80009-x. PMID: 7446550.
6. Ozdemir S, Aydogan O, Koksal Cakirlar F. Biofilm Formation and Antimicrobial Susceptibility of Non-Diphtheria Corynebacterium Strains Isolated from Blood Cultures: First Report from Turkey. Medeni Med J. 2021;36(2):123-129. doi: 10.5222/MMJ.2021.60252. Epub 2021 Jun 18. PMID: 34239764; PMCID: PMC8226407.
7. Tauch A, Kaiser O, Hain T, Goesmann A, Weisshaar B, Albersmeier A, Bekel T, Bischoff N, Brune I, Chakraborty T, Kalinowski J, Meyer F, Rupp O, Schneiker S, Viehoever P, Pühler A. Complete genome sequence and analysis of the multiresistant nosocomial pathogen Corynebacterium jeikeium K411, a lipid-requiring bacterium of the human skin flora. J Bacteriol. 2005 Jul;187(13):4671-82. doi: 10.1128/JB.187.13.4671-4682.2005. PMID: 15968079; PMCID: PMC1151758.

-Arooj Devi MD is a second-year pathology (AP/CP) resident at Brody School of Medicine at East Carolina University. She is interested in breast and gynecological pathology.

-John E. Markantonis DO D(ABMM) is the head of microbiology at ECU Health Medical Center and an assistant professor at Brody School of Medicine at East Carolina University.